DNA replication
In molecular biology,[1][2][3] DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule.[4] DNA replication occurs in all living organisms acting as the most essential part of biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA.[5] The cell possesses the distinctive property of division, which makes replication of DNA essential.
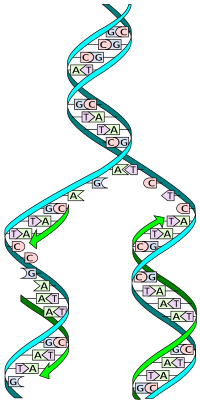
DNA is made up of a double helix of two complementary strands. The double helix describes the appearance of a double-stranded DNA which is thus composed of two linear strands that run opposite to each other and twist together.[6] During replication, these strands are separated. Each strand of the original DNA molecule then serves as a template for the production of its counterpart, a process referred to as semiconservative replication. As a result of semi-conservative replication, the new helix will be composed of an original DNA strand as well as a newly synthesized strand.[7] Cellular proofreading and error-checking mechanisms ensure near perfect fidelity for DNA replication.[8][9]
In a cell, DNA replication begins at specific locations, or origins of replication,[10] in the genome[11] which contains the genetic material of an organism.[12] Unwinding of DNA at the origin and synthesis of new strands, accommodated by an enzyme known as helicase, results in replication forks growing bi-directionally from the origin. A number of proteins are associated with the replication fork to help in the initiation and continuation of DNA synthesis. Most prominently, DNA polymerase synthesizes the new strands by adding nucleotides that complement each (template) strand. DNA replication occurs during the S-stage of interphase.[13]
DNA replication (DNA amplification) can also be performed in vitro (artificially, outside a cell).[14] DNA polymerases isolated from cells and artificial DNA primers can be used to start DNA synthesis at known sequences in a template DNA molecule. Polymerase chain reaction (PCR), ligase chain reaction (LCR), and transcription-mediated amplification (TMA) are examples. In March 2021, researchers reported evidence suggesting that a preliminary form of transfer RNA, a necessary component of translation, the biological synthesis of new proteins in accordance with the genetic code, could have been a replicator molecule itself in the very early development of life, or abiogenesis.[15][16]
DNA structure
[edit]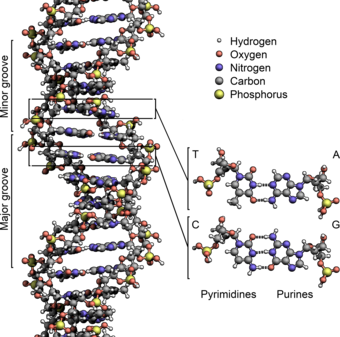
DNA exists as a double-stranded structure, with both strands coiled together to form the characteristic double helix. Each single strand of DNA is a chain of four types of nucleotides. Nucleotides in DNA contain a deoxyribose sugar, a phosphate, and a nucleobase. The four types of nucleotide correspond to the four nucleobases adenine, cytosine, guanine, and thymine, commonly abbreviated as A, C, G, and T. Adenine and guanine are purine[17] bases, while cytosine and thymine are pyrimidines. These nucleotides form phosphodiester bonds, creating the phosphate-deoxyribose backbone of the DNA double helix with the nucleobases pointing inward (i.e., toward the opposing strand). Nucleobases are matched between strands through hydrogen bonds to form base pairs. Adenine pairs with thymine (two hydrogen bonds), and guanine pairs with cytosine (three hydrogen bonds).[18]
DNA strands have a directionality, and the different ends of a single strand are called the "3′ (three-prime) end" and the "5′ (five-prime) end". By convention, if the base sequence of a single strand of DNA is given, the left end of the sequence is the 5′ end, while the right end of the sequence is the 3′ end. The strands of the double helix are anti-parallel, with one being 5′ to 3′, and the opposite strand 3′ to 5′. These terms refer to the carbon atom in deoxyribose to which the next phosphate in the chain attaches. Directionality has consequences in DNA synthesis, because DNA polymerase can synthesize DNA in only one direction by adding nucleotides to the 3′ end of a DNA strand.[citation needed]
The pairing of complementary bases in DNA (through hydrogen bonding) means that the information contained within each strand is redundant. Phosphodiester (intra-strand) bonds are stronger than hydrogen (inter-strand) bonds. The actual job of the phosphodiester bonds is where in DNA polymers connect the 5' carbon atom of one nucleotide to the 3' carbon atom of another nucleotide, while the hydrogen bonds stabilize DNA double helices across the helix axis but not in the direction of the axis.[19] This makes it possible to separate the strands from one another. The nucleotides on a single strand can therefore be used to reconstruct nucleotides on a newly synthesized partner strand.[20]
DNA polymerase
[edit]
DNA polymerases are a family of enzymes that carry out all forms of DNA replication.[22] DNA polymerases in general cannot initiate synthesis of new strands but can only extend an existing DNA or RNA strand paired with a template strand. To begin synthesis, a short fragment of RNA, called a primer, must be created and paired with the template DNA strand.
DNA polymerase adds a new strand of DNA by extending the 3′ end of an existing nucleotide chain, adding new nucleotides matched to the template strand, one at a time, via the creation of phosphodiester bonds. The energy for this process of DNA polymerization comes from hydrolysis of the high-energy phosphate (phosphoanhydride) bonds between the three phosphates attached to each unincorporated base. Free bases with their attached phosphate groups are called nucleotides; in particular, bases with three attached phosphate groups are called nucleoside triphosphates. When a nucleotide is being added to a growing DNA strand, the formation of a phosphodiester bond between the proximal phosphate of the nucleotide to the growing chain is accompanied by hydrolysis of a high-energy phosphate bond with release of the two distal phosphate groups as a pyrophosphate. Enzymatic hydrolysis of the resulting pyrophosphate into inorganic phosphate consumes a second high-energy phosphate bond and renders the reaction effectively irreversible.[Note 1]
In general, DNA polymerases are highly accurate, with an intrinsic error rate of less than one mistake for every 107 nucleotides added.[23] Some DNA polymerases can also delete nucleotides from the end of a developing strand in order to fix mismatched bases. This is known as proofreading. Finally, post-replication mismatch repair mechanisms monitor the DNA for errors, being capable of distinguishing mismatches in the newly synthesized DNA Strand from the original strand sequence. Together, these three discrimination steps enable replication fidelity of less than one mistake for every 109 nucleotides added.[23]
The rate of DNA replication in a living cell was first measured as the rate of phage T4 DNA elongation in phage-infected E. coli.[24] During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second. The mutation rate per base pair per replication during phase T4 DNA synthesis is 1.7 per 108.[25]
Replication process
[edit]
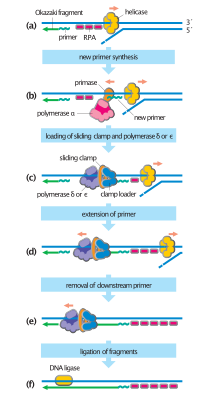
DNA replication, like all biological polymerization processes, proceeds in three enzymatically catalyzed and coordinated steps: initiation, elongation and termination.
Initiation
[edit]

For a cell to divide, it must first replicate its DNA.[26] DNA replication is an all-or-none process; once replication begins, it proceeds to completion. Once replication is complete, it does not occur again in the same cell cycle. This is made possible by the division of initiation of the pre-replication complex.[citation needed]
Pre-replication complex
[edit]In late mitosis and early G1 phase, a large complex of initiator proteins assembles into the pre-replication complex at particular points in the DNA, known as "origins".[11][10] In E. coli the primary initiator protein is Dna A; in yeast, this is the origin recognition complex.[27] Sequences used by initiator proteins tend to be "AT-rich" (rich in adenine and thymine bases), because A-T base pairs have two hydrogen bonds (rather than the three formed in a C-G pair) and thus are easier to strand-separate.[28] In eukaryotes, the origin recognition complex catalyzes the assembly of initiator proteins into the pre-replication complex. In addition, a recent report suggests that budding yeast ORC dimerizes in a cell cycle dependent manner to control licensing.[29][30] In turn, the process of ORC dimerization is mediated by a cell cycle-dependent Noc3p dimerization cycle in vivo, and this role of Noc3p is separable from its role in ribosome biogenesis. An essential Noc3p dimerization cycle mediates ORC double-hexamer formation in replication licensing ORC and Noc3p are continuously bound to the chromatin throughout the cell cycle.[31] Cdc6 and Cdt1 then associate with the bound origin recognition complex at the origin in order to form a larger complex necessary to load the Mcm complex onto the DNA. In eukaryotes, the Mcm complex is the helicase that will split the DNA helix at the replication forks and origins. The Mcm complex is recruited at late G1 phase and loaded by the ORC-Cdc6-Cdt1 complex onto the DNA via ATP-dependent protein remodeling. The loading of the MCM complex onto the origin DNA marks the completion of pre-replication complex formation.[32]
If environmental conditions are right in late G1 phase, the G1 and G1/S cyclin-Cdk complexes are activated, which stimulate expression of genes that encode components of the DNA synthetic machinery. G1/S-Cdk activation also promotes the expression and activation of S-Cdk complexes, which may play a role in activating replication origins depending on species and cell type. Control of these Cdks vary depending on cell type and stage of development. This regulation is best understood in budding yeast, where the S cyclins Clb5 and Clb6 are primarily responsible for DNA replication.[33] Clb5,6-Cdk1 complexes directly trigger the activation of replication origins and are therefore required throughout S phase to directly activate each origin.[32]
In a similar manner, Cdc7 is also required through S phase to activate replication origins. Cdc7 is not active throughout the cell cycle, and its activation is strictly timed to avoid premature initiation of DNA replication. In late G1, Cdc7 activity rises abruptly as a result of association with the regulatory subunit DBF4, which binds Cdc7 directly and promotes its protein kinase activity. Cdc7 has been found to be a rate-limiting regulator of origin activity. Together, the G1/S-Cdks and/or S-Cdks and Cdc7 collaborate to directly activate the replication origins, leading to initiation of DNA synthesis.[32]
Preinitiation complex
[edit]In early S phase, S-Cdk and Cdc7 activation lead to the assembly of the preinitiation complex, a massive protein complex formed at the origin. Formation of the preinitiation complex displaces Cdc6 and Cdt1 from the origin replication complex, inactivating and disassembling the pre-replication complex. Loading the preinitiation complex onto the origin activates the Mcm helicase, causing unwinding of the DNA helix. The preinitiation complex also loads α-primase and other DNA polymerases onto the DNA.[32]
After α-primase synthesizes the first primers, the primer-template junctions interact with the clamp loader, which loads the sliding clamp onto the DNA to begin DNA synthesis. The components of the preinitiation complex remain associated with replication forks as they move out from the origin.[32]
Elongation
[edit]DNA polymerase has 5′–3′ activity. All known DNA replication systems require a free 3′ hydroxyl group before synthesis can be initiated (note: the DNA template is read in 3′ to 5′ direction whereas a new strand is synthesized in the 5′ to 3′ direction—this is often confused). Four distinct mechanisms for DNA synthesis are recognized:[citation needed]
- All cellular life forms and many DNA viruses, phages and plasmids use a primase to synthesize a short RNA primer with a free 3′ OH group which is subsequently elongated by a DNA polymerase.
- The retroelements (including retroviruses) employ a transfer RNA that primes DNA replication by providing a free 3′ OH that is used for elongation by the reverse transcriptase.
- In the adenoviruses and the φ29 family of bacteriophages, the 3′ OH group is provided by the side chain of an amino acid of the genome attached protein (the terminal protein) to which nucleotides are added by the DNA polymerase to form a new strand.
- In the single stranded DNA viruses—a group that includes the circoviruses, the geminiviruses, the parvoviruses and others—and also the many phages and plasmids that use the rolling circle replication (RCR) mechanism, the RCR endonuclease creates a nick in the genome strand (single stranded viruses) or one of the DNA strands (plasmids). The 5′ end of the nicked strand is transferred to a tyrosine residue on the nuclease and the free 3′ OH group is then used by the DNA polymerase to synthesize the new strand.
Cellular organisms use the first of these pathways since it is the most well-known. In this mechanism, once the two strands are separated, primase adds RNA primers to the template strands. The leading strand receives one RNA primer while the lagging strand receives several. The leading strand is continuously extended from the primer by a DNA polymerase with high processivity, while the lagging strand is extended discontinuously from each primer forming Okazaki fragments. RNase removes the primer RNA fragments, and a low processivity DNA polymerase distinct from the replicative polymerase enters to fill the gaps. When this is complete, a single nick on the leading strand and several nicks on the lagging strand can be found. Ligase works to fill these nicks in, thus completing the newly replicated DNA molecule.[citation needed]
The primase used in this process differs significantly between bacteria and archaea/eukaryotes. Bacteria use a primase belonging to the DnaG protein superfamily which contains a catalytic domain of the TOPRIM fold type.[34] The TOPRIM fold contains an α/β core with four conserved strands in a Rossmann-like topology. This structure is also found in the catalytic domains of topoisomerase Ia, topoisomerase II, the OLD-family nucleases and DNA repair proteins related to the RecR protein.[citation needed]
The primase used by archaea and eukaryotes, in contrast, contains a highly derived version of the RNA recognition motif (RRM). This primase is structurally similar to many viral RNA-dependent RNA polymerases, reverse transcriptases, cyclic nucleotide generating cyclases and DNA polymerases of the A/B/Y families that are involved in DNA replication and repair. In eukaryotic replication, the primase forms a complex with Pol α.[35]
Multiple DNA polymerases take on different roles in the DNA replication process. In E. coli, DNA Pol III is the polymerase enzyme primarily responsible for DNA replication. It assembles into a replication complex at the replication fork that exhibits extremely high processivity, remaining intact for the entire replication cycle. In contrast, DNA Pol I is the enzyme responsible for replacing RNA primers with DNA. DNA Pol I has a 5′ to 3′ exonuclease activity in addition to its polymerase activity, and uses its exonuclease activity to degrade the RNA primers ahead of it as it extends the DNA strand behind it, in a process called nick translation. Pol I is much less processive than Pol III because its primary function in DNA replication is to create many short DNA regions rather than a few very long regions.[citation needed]
In eukaryotes, the low-processivity enzyme, Pol α, helps to initiate replication because it forms a complex with primase.[36] In eukaryotes, leading strand synthesis is thought to be conducted by Pol ε; however, this view has recently been challenged, suggesting a role for Pol δ.[37] Primer removal is completed Pol δ[38] while repair of DNA during replication is completed by Pol ε.
As DNA synthesis continues, the original DNA strands continue to unwind on each side of the bubble, forming a replication fork with two prongs. In bacteria, which have a single origin of replication on their circular chromosome, this process creates a "theta structure" (resembling the Greek letter theta: θ). In contrast, eukaryotes have longer linear chromosomes and initiate replication at multiple origins within these.[39]
Replication fork
[edit]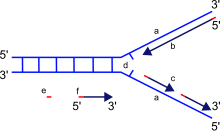
a: template, b: leading strand, c: lagging strand, d: replication fork, e: primer, f: Okazaki fragments

The replication fork is a structure that forms within the long helical DNA during DNA replication. It is produced by enzymes called helicases that break the hydrogen bonds that hold the DNA strands together in a helix. The resulting structure has two branching "prongs", each one made up of a single strand of DNA. These two strands serve as the template for the leading and lagging strands, which will be created as DNA polymerase matches complementary nucleotides to the templates; the templates may be properly referred to as the leading strand template and the lagging strand template.[citation needed]
DNA is read by DNA polymerase in the 3′ to 5′ direction, meaning the new strand is synthesized in the 5' to 3' direction. Since the leading and lagging strand templates are oriented in opposite directions at the replication fork, a major issue is how to achieve synthesis of new lagging strand DNA, whose direction of synthesis is opposite to the direction of the growing replication fork.[citation needed]
Leading strand
[edit]The leading strand is the strand of new DNA which is synthesized in the same direction as the growing replication fork. This sort of DNA replication is continuous.[citation needed]
Lagging strand
[edit]The lagging strand is the strand of new DNA whose direction of synthesis is opposite to the direction of the growing replication fork. Because of its orientation, replication of the lagging strand is more complicated as compared to that of the leading strand. As a consequence, the DNA polymerase on this strand is seen to "lag behind" the other strand.[citation needed]
The lagging strand is synthesized in short, separated segments. On the lagging strand template, a primase "reads" the template DNA and initiates synthesis of a short complementary RNA primer. A DNA polymerase extends the primed segments, forming Okazaki fragments. The RNA primers are then removed and replaced with DNA, and the fragments of DNA are joined by DNA ligase.[citation needed]
Dynamics at the replication fork
[edit]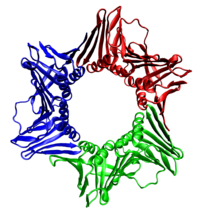
In all cases the helicase is composed of six polypeptides that wrap around only one strand of the DNA being replicated. The two polymerases are bound to the helicase hexamer. In eukaryotes the helicase wraps around the leading strand, and in prokaryotes it wraps around the lagging strand.[40]
As helicase unwinds DNA at the replication fork, the DNA ahead is forced to rotate. This process results in a build-up of twists in the DNA ahead.[41] This build-up creates a torsional load that would eventually stop the replication fork. Topoisomerases are enzymes that temporarily break the strands of DNA, relieving the tension caused by unwinding the two strands of the DNA helix; topoisomerases (including DNA gyrase) achieve this by adding negative supercoils to the DNA helix.[42]
Bare single-stranded DNA tends to fold back on itself forming secondary structures; these structures can interfere with the movement of DNA polymerase. To prevent this, single-strand binding proteins bind to the DNA until a second strand is synthesized, preventing secondary structure formation.[43]
Double-stranded DNA is coiled around histones that play an important role in regulating gene expression so the replicated DNA must be coiled around histones at the same places as the original DNA.[44] To ensure this, histone chaperones disassemble the chromatin before it is replicated and replace the histones in the correct place. Some steps in this reassembly are somewhat speculative.[45]
Clamp proteins act as a sliding clamp on DNA, allowing the DNA polymerase to bind to its template and aid in processivity. The inner face of the clamp enables DNA to be threaded through it. Once the polymerase reaches the end of the template or detects double-stranded DNA, the sliding clamp undergoes a conformational change that releases the DNA polymerase. Clamp-loading proteins are used to initially load the clamp, recognizing the junction between template and RNA primers.[9]:274-5
DNA replication proteins
[edit]At the replication fork, many replication enzymes assemble on the DNA into a complex molecular machine called the replisome. The following is a list of major DNA replication enzymes that participate in the replisome:[46]
| Enzyme | Function in DNA replication |
|---|---|
| DNA helicase | Also known as helix destabilizing enzyme. Helicase separates the two strands of DNA at the Replication Fork behind the topoisomerase. |
| DNA polymerase | The enzyme responsible for catalyzing the addition of nucleotide substrates to DNA in the 5′ to 3′ direction during DNA replication. Also performs proof-reading and error correction. There exist many different types of DNA Polymerase, each of which perform different functions in different types of cells. |
| DNA clamp | A protein which prevents elongating DNA polymerases from dissociating from the DNA parent strand. |
| Single-strand DNA-binding protein | Bind to ssDNA and prevent the DNA double helix from re-annealing after DNA helicase unwinds it, thus maintaining the strand separation, and facilitating the synthesis of the new strand. |
| Topoisomerase | Relaxes the DNA from its super-coiled nature. |
| DNA gyrase | Relieves strain of unwinding by DNA helicase; this is a specific type of topoisomerase |
| DNA ligase | Re-anneals the semi-conservative strands and joins Okazaki Fragments of the lagging strand. |
| Primase | Provides a starting point of RNA (or DNA) for DNA polymerase to begin synthesis of the new DNA strand. |
| Telomerase | Lengthens telomeric DNA by adding repetitive nucleotide sequences to the ends of eukaryotic chromosomes. This allows germ cells and stem cells to avoid the Hayflick limit on cell division.[47] |
In vitro single-molecule experiments (using optical tweezers and magnetic tweezers) have found synergetic interactions between the replisome enzymes (helicase, polymerase, and Single-strand DNA-binding protein) and with the DNA replication fork enhancing DNA-unwinding and DNA-replication.[14] These results lead to the development of kinetic models accounting for the synergetic interactions and their stability.[14]
Replication machinery
[edit]
Replication machineries consist of factors involved in DNA replication and appearing on template ssDNAs. Replication machineries include primosotors are replication enzymes; DNA polymerase, DNA helicases, DNA clamps and DNA topoisomerases, and replication proteins; e.g. single-stranded DNA binding proteins (SSB). In the replication machineries these components coordinate. In most of the bacteria, all of the factors involved in DNA replication are located on replication forks and the complexes stay on the forks during DNA replication. Replication machineries are also referred to as replisomes, or DNA replication systems. These terms are generic terms for proteins located on replication forks. In eukaryotic and some bacterial cells the replisomes are not formed.[citation needed]
In an alternative figure, DNA factories are similar to projectors and DNAs are like as cinematic films passing constantly into the projectors. In the replication factory model, after both DNA helicases for leading strands and lagging strands are loaded on the template DNAs, the helicases run along the DNAs into each other. The helicases remain associated for the remainder of replication process. Peter Meister et al. observed directly replication sites in budding yeast by monitoring green fluorescent protein (GFP)-tagged DNA polymerases α. They detected DNA replication of pairs of the tagged loci spaced apart symmetrically from a replication origin and found that the distance between the pairs decreased markedly by time.[48] This finding suggests that the mechanism of DNA replication goes with DNA factories. That is, couples of replication factories are loaded on replication origins and the factories associated with each other. Also, template DNAs move into the factories, which bring extrusion of the template ssDNAs and new DNAs. Meister's finding is the first direct evidence of replication factory model. Subsequent research has shown that DNA helicases form dimers in many eukaryotic cells and bacterial replication machineries stay in single intranuclear location during DNA synthesis.[49]
Replication Factories Disentangle Sister Chromatids. The disentanglement is essential for distributing the chromatids into daughter cells after DNA replication. Because sister chromatids after DNA replication hold each other by Cohesin rings, there is the only chance for the disentanglement in DNA replication. Fixing of replication machineries as replication factories can improve the success rate of DNA replication. If replication forks move freely in chromosomes, catenation of nuclei is aggravated and impedes mitotic segregation.[48]
Termination
[edit]Eukaryotes initiate DNA replication at multiple points in the chromosome, so replication forks meet and terminate at many points in the chromosome. Because eukaryotes have linear chromosomes, DNA replication is unable to reach the very end of the chromosomes. Due to this problem, DNA is lost in each replication cycle from the end of the chromosome. Telomeres are regions of repetitive DNA close to the ends and help prevent loss of genes due to this shortening. Shortening of the telomeres is a normal process in somatic cells. This shortens the telomeres of the daughter DNA chromosome. As a result, cells can only divide a certain number of times before the DNA loss prevents further division. (This is known as the Hayflick limit.) Within the germ cell line, which passes DNA to the next generation, telomerase extends the repetitive sequences of the telomere region to prevent degradation. Telomerase can become mistakenly active in somatic cells, sometimes leading to cancer formation. Increased telomerase activity is one of the hallmarks of cancer.[citation needed]
Termination requires that the progress of the DNA replication fork must stop or be blocked. Termination at a specific locus, when it occurs, involves the interaction between two components: (1) a termination site sequence in the DNA, and (2) a protein which binds to this sequence to physically stop DNA replication. In various bacterial species, this is named the DNA replication terminus site-binding protein, or Ter protein.[citation needed]
Because bacteria have circular chromosomes, termination of replication occurs when the two replication forks meet each other on the opposite end of the parental chromosome. E. coli regulates this process through the use of termination sequences that, when bound by the Tus protein, enable only one direction of replication fork to pass through. As a result, the replication forks are constrained to always meet within the termination region of the chromosome.[50]
Regulation
[edit]
Eukaryotes
[edit]Within eukaryotes, DNA replication is controlled within the context of the cell cycle. As the cell grows and divides, it progresses through stages in the cell cycle; DNA replication takes place during the S phase (synthesis phase). The progress of the eukaryotic cell through the cycle is controlled by cell cycle checkpoints. Progression through checkpoints is controlled through complex interactions between various proteins, including cyclins and cyclin-dependent kinases.[51] Unlike bacteria, eukaryotic DNA replicates in the confines of the nucleus.[52]
The G1/S checkpoint (restriction checkpoint) regulates whether eukaryotic cells enter the process of DNA replication and subsequent division. Cells that do not proceed through this checkpoint remain in the G0 stage and do not replicate their DNA.[citation needed]
Once the DNA has gone through the "G1/S" test, it can only be copied once in every cell cycle. When the Mcm complex moves away from the origin, the pre-replication complex is dismantled. Because a new Mcm complex cannot be loaded at an origin until the pre-replication subunits are reactivated, one origin of replication can not be used twice in the same cell cycle.[32]
Activation of S-Cdks in early S phase promotes the destruction or inhibition of individual pre-replication complex components, preventing immediate reassembly. S and M-Cdks continue to block pre-replication complex assembly even after S phase is complete, ensuring that assembly cannot occur again until all Cdk activity is reduced in late mitosis.[32]
In budding yeast, inhibition of assembly is caused by Cdk-dependent phosphorylation of pre-replication complex components. At the onset of S phase, phosphorylation of Cdc6 by Cdk1 causes the binding of Cdc6 to the SCF ubiquitin protein ligase, which causes proteolytic destruction of Cdc6. Cdk-dependent phosphorylation of Mcm proteins promotes their export out of the nucleus along with Cdt1 during S phase, preventing the loading of new Mcm complexes at origins during a single cell cycle. Cdk phosphorylation of the origin replication complex also inhibits pre-replication complex assembly. The individual presence of any of these three mechanisms is sufficient to inhibit pre-replication complex assembly. However, mutations of all three proteins in the same cell does trigger reinitiation at many origins of replication within one cell cycle.[32][53]
In animal cells, the protein geminin is a key inhibitor of pre-replication complex assembly. Geminin binds Cdt1, preventing its binding to the origin recognition complex. In G1, levels of geminin are kept low by the APC, which ubiquitinates geminin to target it for degradation. When geminin is destroyed, Cdt1 is released, allowing it to function in pre-replication complex assembly. At the end of G1, the APC is inactivated, allowing geminin to accumulate and bind Cdt1.[32]
Replication of chloroplast and mitochondrial genomes occurs independently of the cell cycle, through the process of D-loop replication.[citation needed]
Replication focus
[edit]In vertebrate cells, replication sites concentrate into positions called replication foci.[48] Replication sites can be detected by immunostaining daughter strands and replication enzymes and monitoring GFP-tagged replication factors. By these methods it is found that replication foci of varying size and positions appear in S phase of cell division and their number per nucleus is far smaller than the number of genomic replication forks.
P. Heun et al.,[48](2001) tracked GFP-tagged replication foci in budding yeast cells and revealed that replication origins move constantly in G1 and S phase and the dynamics decreased significantly in S phase.[48] Traditionally, replication sites were fixed on spatial structure of chromosomes by nuclear matrix or lamins. The Heun's results denied the traditional concepts, budding yeasts do not have lamins, and support that replication origins self-assemble and form replication foci.[citation needed]
By firing of replication origins, controlled spatially and temporally, the formation of replication foci is regulated. D. A. Jackson et al.(1998) revealed that neighboring origins fire simultaneously in mammalian cells.[48] Spatial juxtaposition of replication sites brings clustering of replication forks. The clustering do rescue of stalled replication forks and favors normal progress of replication forks. Progress of replication forks is inhibited by many factors; collision with proteins or with complexes binding strongly on DNA, deficiency of dNTPs, nicks on template DNAs and so on. If replication forks get stuck and the rest of the sequences from the stuck forks are not copied, then the daughter strands get nick nick unreplicated sites. The un-replicated sites on one parent's strand hold the other strand together but not daughter strands. Therefore, the resulting sister chromatids cannot separate from each other and cannot divide into 2 daughter cells. When neighboring origins fire and a fork from one origin is stalled, fork from other origin access on an opposite direction of the stalled fork and duplicate the un-replicated sites. As other mechanism of the rescue there is application of dormant replication origins that excess origins do not fire in normal DNA replication.[citation needed]
Bacteria
[edit]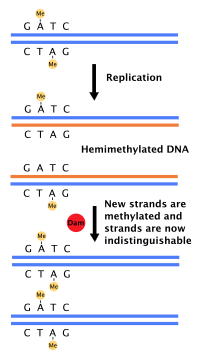
Most bacteria do not go through a well-defined cell cycle but instead continuously copy their DNA; during rapid growth, this can result in the concurrent occurrence of multiple rounds of replication.[54] In E. coli, the best-characterized bacteria, DNA replication is regulated through several mechanisms, including: the hemimethylation and sequestering of the origin sequence, the ratio of adenosine triphosphate (ATP) to adenosine diphosphate (ADP), and the levels of protein DnaA. All these control the binding of initiator proteins to the origin sequences.[55]
Because E. coli methylates GATC DNA sequences, DNA synthesis results in hemimethylated sequences. This hemimethylated DNA is recognized by the protein SeqA, which binds and sequesters the origin sequence; in addition, DnaA (required for initiation of replication) binds less well to hemimethylated DNA. As a result, newly replicated origins are prevented from immediately initiating another round of DNA replication.[56]
ATP builds up when the cell is in a rich medium, triggering DNA replication once the cell has reached a specific size. ATP competes with ADP to bind to DnaA, and the DnaA-ATP complex is able to initiate replication. A certain number of DnaA proteins are also required for DNA replication — each time the origin is copied, the number of binding sites for DnaA doubles, requiring the synthesis of more DnaA to enable another initiation of replication.[citation needed]
In fast-growing bacteria, such as E. coli, chromosome replication takes more time than dividing the cell. The bacteria solve this by initiating a new round of replication before the previous one has been terminated.[57] The new round of replication will form the chromosome of the cell that is born two generations after the dividing cell. This mechanism creates overlapping replication cycles.
Problems with DNA replication
[edit]This section needs expansion. You can help by adding to it. (May 2020) |
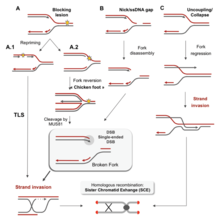
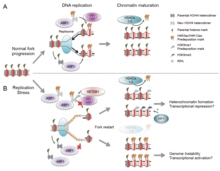
There are many events that contribute to replication stress, including:[58]
- Misincorporation of ribonucleotides
- Unusual DNA structures
- Conflicts between replication and transcription
- Insufficiency of essential replication factors
- Common fragile sites
- Overexpression or constitutive activation of oncogenes
- Chromatin inaccessibility
Polymerase chain reaction
[edit]Researchers commonly replicate DNA in vitro using the polymerase chain reaction (PCR). PCR uses a pair of primers to span a target region in template DNA, and then polymerizes partner strands in each direction from these primers using a thermostable DNA polymerase. Repeating this process through multiple cycles amplifies the targeted DNA region. At the start of each cycle, the mixture of template and primers is heated, separating the newly synthesized molecule and template. Then, as the mixture cools, both of these become templates for annealing of new primers, and the polymerase extends from these. As a result, the number of copies of the target region doubles each round, increasing exponentially.[59]
See also
[edit]- Autopoiesis
- Cell (biology)
- Cell division
- Chromosome segregation
- Data storage device
- Gene
- Gene expression
- Epigenetics
- Genome
- Hachimoji DNA
- Life
- Replication (computing)
- Self-replication
Notes
[edit]- ^ The energetics of this process may also help explain the directionality of synthesis—if DNA were synthesized in the 3′ to 5′ direction, the energy for the process would come from the 5′ end of the growing strand rather than from free nucleotides. The problem is that if the high energy triphosphates were on the growing strand and not on the free nucleotides, proof-reading by removing a mismatched terminal nucleotide would be problematic: Once a nucleotide is added, the triphosphate is lost and a single phosphate remains on the backbone between the new nucleotide and the rest of the strand. If the added nucleotide were mismatched, removal would result in a DNA strand terminated by a monophosphate at the end of the "growing strand" rather than a high energy triphosphate. So strand would be stuck and wouldn't be able to grow anymore. In actuality, the high energy triphosphates hydrolyzed at each step originate from the free nucleotides, not the polymerized strand, so this issue does not exist.
References
[edit]- ^ O'Donnell M, Langston L, Stillman B (July 2013). "Principles and concepts of DNA replication in bacteria, archaea, and eukarya". Cold Spring Harbor Perspectives in Biology. 5 (7): a010108. doi:10.1101/cshperspect.a010108. PMC 3685895. PMID 23818497.
- ^ Prioleau MN, MacAlpine DM (August 2016). "DNA replication origins-where do we begin?". Genes & Development. 30 (15): 1683–1697. doi:10.1101/gad.285114.116. PMC 5002974. PMID 27542827.
- ^ Dewar JM, Walter JC (August 2017). "Mechanisms of DNA replication termination". Nature Reviews. Molecular Cell Biology. 18 (8): 507–516. doi:10.1038/nrm.2017.42. PMC 6386472. PMID 28537574.
- ^ Sabhadiya A (2022-03-01). "What Is DNA Replication And Its Steps?". Retrieved 2023-08-04.
- ^ "GENETICS / DNA REPLICATION (BASIC) – Pathwayz". pathwayz.org. Retrieved 2020-12-10.
- ^ "double helix". Learn Science at Scitable. Nature Education. Retrieved 2020-12-10.
- ^ Pray LA (2008). "Semi-Conservative DNA Replication; Meselson and Stahl". Nature Education. 1 (1): 98.
- ^ Imperfect DNA replication results in mutations. Berg JM, Tymoczko JL, Stryer L, Clarke ND (2002). "Chapter 27: DNA Replication, Recombination, and Repair". Biochemistry. W.H. Freeman and Company. ISBN 0-7167-3051-0. Archived from the original on 2020-03-26. Retrieved 2019-08-09.
- ^ a b Lodish H, Berk A, Zipursky SL, et al. (2000). "DNA Replication, Repair, and Recombination". Molecular Cell Biology (4th ed.). WH Freeman. ISBN 0-7167-3136-3.
- ^ a b Hu Y, Stillman B (February 2023). "Origins of DNA replication in eukaryotes". Molecular Cell. 83 (3): 352–372. doi:10.1016/j.molcel.2022.12.024. PMC 9898300. PMID 36640769.
- ^ a b Berg JM, Tymoczko JL, Stryer L, Clarke ND (2002). "Chapter 27, Section 4: DNA Replication of Both Strands Proceeds Rapidly from Specific Start Sites". Biochemistry. W.H. Freeman and Company. ISBN 0-7167-3051-0. Archived from the original on 2020-03-26. Retrieved 2019-08-09.
- ^ "What is a genome?". yourgenome. Retrieved 2020-12-10.
- ^ Chagin, Vadim O.; Stear, Jeffrey H.; Cardoso, M. Cristina (April 2010). "Organization of DNA replication". Cold Spring Harbor Perspectives in Biology. 2 (4): a000737. doi:10.1101/cshperspect.a000737. ISSN 1943-0264. PMC 2845211. PMID 20452942.
- ^ a b c Jarillo J, Ibarra B, Cao-García FJ (2021). "DNA replication: In vitro single-molecule manipulation data analysis and models". Computational and Structural Biotechnology Journal. 19: 3765–3778. doi:10.1016/j.csbj.2021.06.032. PMC 8267548. PMID 34285777.
- ^ Kühnlein A, Lanzmich SA, Braun D (March 2021). "tRNA sequences can assemble into a replicator". eLife. 10: e63431. doi:10.7554/eLife.63431. PMC 7924937. PMID 33648631.
- ^ Maximilian L (3 April 2021). "Solving the Chicken-and-the-Egg Problem – "A Step Closer to the Reconstruction of the Origin of Life"". SciTechDaily. Retrieved 3 April 2021.
- ^ Friedberg EC (January 2003). "DNA damage and repair". Nature. 421 (6921): 436–440. Bibcode:2003Natur.421..436F. doi:10.1038/nature01408. PMID 12540918.
- ^ Sabhadiya A (2023-03-13). "Base Pair: Definition, Rules In DNA And RNA". Retrieved 2023-08-04.
- ^ "DNA function & structure (with diagram) (article)". Khan Academy. Retrieved 2020-12-10.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). Molecular Biology of the Cell (4th ed.). Garland Science. pp. 238–240. ISBN 0-8153-3218-1.
- ^ Allison L (2007). Fundamental Molecular Biology. Blackwell Publishing. p. 112. ISBN 978-1-4051-0379-4.
- ^ Berg JM, Tymoczko JL, Stryer L, Clarke ND (2002). Biochemistry. W.H. Freeman and Company. ISBN 0-7167-3051-0. Chapter 27, Section 2: DNA Polymerases Require a Template and a Primer
- ^ a b McCulloch SD, Kunkel TA (January 2008). "The fidelity of DNA synthesis by eukaryotic replicative and translesion synthesis polymerases". Cell Research. 18 (1): 148–161. doi:10.1038/cr.2008.4. PMC 3639319. PMID 18166979.
- ^ McCarthy D, Minner C, Bernstein H, Bernstein C (October 1976). "DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant". Journal of Molecular Biology. 106 (4): 963–981. doi:10.1016/0022-2836(76)90346-6. PMID 789903.
- ^ Drake JW (1970) The Molecular Basis of Mutation. Holden-Day, San Francisco ISBN 0-8162-2450-1 ISBN 978-0-8162-2450-0
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "Chapter 5: DNA Replication Mechanisms". Molecular Biology of the Cell. Garland Science. ISBN 0-8153-3218-1.
- ^ Weigel C, Schmidt A, Rückert B, Lurz R, Messer W (November 1997). "DnaA protein binding to individual DnaA boxes in the Escherichia coli replication origin, oriC". The EMBO Journal. 16 (21): 6574–6583. doi:10.1093/emboj/16.21.6574. PMC 1170261. PMID 9351837.
- ^ Lodish H, Berk A, Zipursky LS, Matsudaira P, Baltimore D, Darnell J (2000). Molecular Cell Biology. W. H. Freeman and Company. ISBN 0-7167-3136-3.12.1. General Features of Chromosomal Replication: Three Common Features of Replication Origins
- ^ Lin YC, Prasanth SG (July 2021). "Replication initiation: Implications in genome integrity". DNA Repair. 103: 103131. doi:10.1016/j.dnarep.2021.103131. PMC 8296962. PMID 33992866.
- ^ Amin A, Wu R, Cheung MH, Scott JF, Wang Z, Zhou Z, et al. (March 2020). "An Essential and Cell-Cycle-Dependent ORC Dimerization Cycle Regulates Eukaryotic Chromosomal DNA Replication". Cell Reports. 30 (10): 3323–3338.e6. doi:10.1016/j.celrep.2020.02.046. PMID 32160540.
- ^ Zhang Y, Yu Z, Fu X, Liang C (June 2002). "Noc3p, a bHLH protein, plays an integral role in the initiation of DNA replication in budding yeast". Cell. 109 (7): 849–860. doi:10.1016/s0092-8674(02)00805-x. PMID 12110182.
- ^ a b c d e f g h i Morgan DO (2007). The cell cycle: principles of control. London: New Science Press. pp. 64–75. ISBN 978-0-19-920610-0. OCLC 70173205.
- ^ Donaldson AD, Raghuraman MK, Friedman KL, Cross FR, Brewer BJ, Fangman WL (August 1998). "CLB5-dependent activation of late replication origins in S. cerevisiae". Molecular Cell. 2 (2): 173–182. doi:10.1016/s1097-2765(00)80127-6. PMID 9734354.
- ^ Aravind L, Leipe DD, Koonin EV (September 1998). "Toprim--a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins". Nucleic Acids Research. 26 (18): 4205–4213. doi:10.1093/nar/26.18.4205. PMC 147817. PMID 9722641.
- ^ Frick DN, Richardson CC (July 2001). "DNA primases". Annual Review of Biochemistry. 70: 39–80. doi:10.1146/annurev.biochem.70.1.39. PMID 11395402. S2CID 33197061.
- ^ Barry ER, Bell SD (December 2006). "DNA replication in the archaea". Microbiology and Molecular Biology Reviews. 70 (4): 876–887. doi:10.1128/MMBR.00029-06. PMC 1698513. PMID 17158702.
- ^ Stillman B (July 2015). "Reconsidering DNA Polymerases at the Replication Fork in Eukaryotes". Molecular Cell. 59 (2): 139–141. doi:10.1016/j.molcel.2015.07.004. PMC 4636199. PMID 26186286.
- ^ Rossi ML (February 2009). Distinguishing the pathways of primer removal during Eukaryotic Okazaki fragment maturation (Ph.D. thesis). School of Medicine and Dentistry, University of Rochester. hdl:1802/6537.
- ^ Huberman JA, Riggs AD (March 1968). "On the mechanism of DNA replication in mammalian chromosomes". Journal of Molecular Biology. 32 (2): 327–341. doi:10.1016/0022-2836(68)90013-2. PMID 5689363.
- ^ Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, et al. (February 2019). "Structures and operating principles of the replisome". Science. 363 (6429): 835. doi:10.1126/science.aav7003. PMC 6681829. PMID 30679383.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "DNA Replication Mechanisms: DNA Topoisomerases Prevent DNA Tangling During Replication". Molecular Biology of the Cell. Garland Science. ISBN 0-8153-3218-1.
- ^ Reece RJ, Maxwell A (26 September 2008). "DNA gyrase: structure and function". Critical Reviews in Biochemistry and Molecular Biology. 26 (3–4): 335–375. doi:10.3109/10409239109114072. PMID 1657531.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "DNA Replication Mechanisms: Special Proteins Help to Open Up the DNA Double Helix in Front of the Replication Fork". Molecular Biology of the Cell. Garland Science. ISBN 0-8153-3218-1.
- ^ Koonin, Eugene V.; Krupovic, Mart; Ishino, Sonoko; Ishino, Yoshizumi (2020-06-09). "The replication machinery of LUCA: common origin of DNA replication and transcription". BMC Biology. 18 (1): 61. doi:10.1186/s12915-020-00800-9. ISSN 1741-7007. PMC 7281927. PMID 32517760.
- ^ Ransom M, Dennehey BK, Tyler JK (January 2010). "Chaperoning histones during DNA replication and repair". Cell. 140 (2): 183–195. doi:10.1016/j.cell.2010.01.004. PMC 3433953. PMID 20141833.
- ^ Griffiths AJ, Wessler SR, Lewontin RC, Carroll SB (2008). Introduction to Genetic Analysis. W. H. Freeman and Company. ISBN 978-0-7167-6887-6.[Chapter 7: DNA: Structure and Replication. pg 283–290]
- ^ Clark J (2009-05-11). "Will the Hayflick limit keep us from living forever?". Howstuffworks. Retrieved January 20, 2015.
- ^ a b c d e f Meister P, Taddei A, Gasser SM (June 2006). "In and out of the replication factory". Cell. 125 (7): 1233–5. doi:10.1016/j.cell.2006.06.014. PMID 16814710. S2CID 15397410.
- ^ Watson JD, Baker TA, Bell SP, Gann A, Levine M, Losick R, Inglis CH (2008). Molecular Biology of the Gene (6th ed.). San Francisco: Pearson/Benjamin Cummings. p. 237. ISBN 978-0-8053-9592-1.
- ^ Brown TA (2002). "Chapter 13.2.3. Termination of replication". Genomes. BIOS Scientific Publishers. ISBN 1-85996-228-9.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "Intracellular Control of Cell-Cycle Events: S-Phase Cyclin-Cdk Complexes (S-Cdks) Initiate DNA Replication Once Per Cycle". Molecular Biology of the Cell. Garland Science. ISBN 0-8153-3218-1.
- ^ Brown TA (2002). "Chapter 13: Genome Replication". Genomes (2nd ed.). Oxford: Wiley-Liss.
- ^ Nguyen VQ, Co C, Li JJ (June 2001). "Cyclin-dependent kinases prevent DNA re-replication through multiple mechanisms". Nature. 411 (6841): 1068–1073. Bibcode:2001Natur.411.1068N. doi:10.1038/35082600. PMID 11429609. S2CID 4393812.
- ^ Tobiason DM, Seifert HS (June 2006). "The obligate human pathogen, Neisseria gonorrhoeae, is polyploid". PLOS Biology. 4 (6): e185. doi:10.1371/journal.pbio.0040185. PMC 1470461. PMID 16719561.
- ^ O'Donnell, Michael; Langston, Lance; Stillman, Bruce (2013-07-01). "Principles and concepts of DNA replication in bacteria, archaea, and eukarya". Cold Spring Harbor Perspectives in Biology. 5 (7): a010108. doi:10.1101/cshperspect.a010108. ISSN 1943-0264. PMC 3685895. PMID 23818497.
- ^ Slater S, Wold S, Lu M, Boye E, Skarstad K, Kleckner N (September 1995). "E. coli SeqA protein binds oriC in two different methyl-modulated reactions appropriate to its roles in DNA replication initiation and origin sequestration". Cell. 82 (6): 927–936. doi:10.1016/0092-8674(95)90272-4. PMID 7553853. S2CID 14652024.
- ^ Cooper S, Helmstetter CE (February 1968). "Chromosome replication and the division cycle of Escherichia coli B/r". Journal of Molecular Biology. 31 (3): 519–540. doi:10.1016/0022-2836(68)90425-7. PMID 4866337.
- ^ Zeman MK, Cimprich KA (January 2014). "Causes and consequences of replication stress". Nature Cell Biology. 16 (1): 2–9. doi:10.1038/ncb2897. PMC 4354890. PMID 24366029.
- ^ Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, et al. (January 1988). "Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase". Science. 239 (4839): 487–491. Bibcode:1988Sci...239..487S. doi:10.1126/science.239.4839.487. PMID 2448875.
