X chromosome
| Human X chromosome | |
|---|---|
Human X chromosome (after G-banding) | |
 X chromosome in human male karyogram | |
| Features | |
| Length (bp) | 154,259,566 bp (CHM13) |
| No. of genes | 804 (CCDS) |
| Type | Allosome |
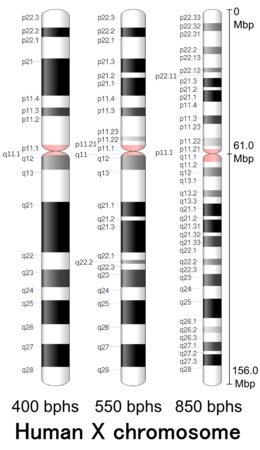
| Centromere position | Submetacentric[1] (61.0 Mbp)[2] |
| Complete gene lists | |
| CCDS | Gene list |
| HGNC | Gene list |
| UniProt | Gene list |
| NCBI | Gene list |
| External map viewers | |
| Ensembl | Chromosome X |
| Entrez | Chromosome X |
| NCBI | Chromosome X |
| UCSC | Chromosome X |
| Full DNA sequences | |
| RefSeq | NC_000023 (FASTA) |
| GenBank | CM000685 (FASTA) |
The X chromosome is one of the two sex chromosomes in many organisms, including mammals, and is found in both males and females. It is a part of the XY sex-determination system and XO sex-determination system. The X chromosome was named for its unique properties by early researchers, which resulted in the naming of its counterpart Y chromosome, for the next letter in the alphabet, following its subsequent discovery.[3]
Discovery
[edit]It was first noted that the X chromosome was special in 1890 by Hermann Henking in Leipzig. Henking was studying the testicles of Pyrrhocoris and noticed that one chromosome did not take part in meiosis. Chromosomes are so named because of their ability to take up staining (chroma in Greek means color). Although the X chromosome could be stained just as well as the others, Henking was unsure whether it was a different class of the object and consequently named it X element,[4] which later became X chromosome after it was established that it was indeed a chromosome.[5]
The idea that the X chromosome was named after its similarity to the letter "X" is mistaken. All chromosomes normally appear as an amorphous blob under the microscope and take on a well-defined shape only during mitosis. This shape is vaguely X-shaped for all chromosomes. It is entirely coincidental that the Y chromosome, during mitosis, has two very short branches which can look merged under the microscope and appear as the descender of a Y-shape.[6]
It was first suggested that the X chromosome was involved in sex determination by Clarence Erwin McClung in 1901. After comparing his work on locusts with Henking's and others, McClung noted that only half the sperm received an X chromosome. He called this chromosome an accessory chromosome, and insisted (correctly) that it was a proper chromosome, and theorized (incorrectly) that it was the male-determining chromosome.[4]
Inheritance pattern
[edit]This section relies largely or entirely upon a single source. (August 2021) |

Luke Hutchison noticed that a number of possible ancestors on the X chromosome inheritance line at a given ancestral generation follows the Fibonacci sequence.[7] A male individual has an X chromosome, which he received from his mother, and a Y chromosome, which he received from his father. The male counts as the "origin" of his own X chromosome (), and at his parents' generation, his X chromosome came from a single parent (). The male's mother received one X chromosome from her mother (the son's maternal grandmother), and one from her father (the son's maternal grandfather), so two grandparents contributed to the male descendant's X chromosome (). The maternal grandfather received his X chromosome from his mother, and the maternal grandmother received X chromosomes from both of her parents, so three great-grandparents contributed to the male descendant's X chromosome (). Five great-great-grandparents contributed to the male descendant's X chromosome (), etc. (Note that this assumes that all ancestors of a given descendant are independent, but if any genealogy is traced far enough back in time, ancestors begin to appear on multiple lines of the genealogy, until eventually, a population founder appears on all lines of the genealogy.)
Humans
[edit]Function
[edit]
The X chromosome in humans spans more than 153 million base pairs (the building material of DNA). It represents about 800 protein-coding genes compared to the Y chromosome containing about 70 genes, out of 20,000–25,000 total genes in the human genome. Each person usually has one pair of sex chromosomes in each cell. Females typically have two X chromosomes, whereas males typically have one X and one Y chromosome. Both males and females retain one of their mother's X chromosomes, and females retain their second X chromosome from their father. Since the father retains his X chromosome from his mother, a human female has one X chromosome from her paternal grandmother (father's side), and one X chromosome from her mother. This inheritance pattern follows the Fibonacci numbers at a given ancestral depth.[citation needed]
Genetic disorders that are due to mutations in genes on the X chromosome are described as X linked. If the X chromosome has a genetic disease gene, it always causes illness in male patients, since men have only one X chromosome and therefore only one copy of each gene. Females, instead, require both X chromosomes to have the illness, and as a result could potentially only be a carrier of genetic illness, since their second X chromosome overrides the first. For example, hemophilia A and B and congenital red–green color blindness run in families this way.
The X chromosome carries hundreds of genes but few, if any, of these have anything to do directly with sex determination. Early in embryonic development in females, one of the two X chromosomes is permanently inactivated in nearly all somatic cells (cells other than egg and sperm cells). This phenomenon is called X-inactivation or Lyonization, and creates a Barr body. If X-inactivation in the somatic cell meant a complete de-functionalizing of one of the X-chromosomes, it would ensure that females, like males, had only one functional copy of the X chromosome in each somatic cell. This was previously assumed to be the case. However, recent research suggests that the Barr body may be more biologically active than was previously supposed.[8]
The partial inactivation of the X-chromosome is due to repressive heterochromatin that compacts the DNA and prevents the expression of most genes. Heterochromatin compaction is regulated by Polycomb Repressive Complex 2 (PRC2).[9]
Genes
[edit]Number of genes
[edit]The following are some of the gene count estimates of human X chromosome. Because researchers use different approaches to genome annotation their predictions of the number of genes on each chromosome varies (for technical details, see gene prediction). Among various projects, the collaborative consensus coding sequence project (CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes.[10]
| Estimated by | Protein-coding genes | Non-coding RNA genes | Pseudogenes | Source | Release date |
|---|---|---|---|---|---|
| CCDS | 804 | — | — | [11] | 2016-09-08 |
| HGNC | 825 | 260 | 606 | [12] | 2017-05-12 |
| Ensembl | 841 | 639 | 871 | [13] | 2017-03-29 |
| UniProt | 839 | — | — | [14] | 2018-02-28 |
| NCBI | 874 | 494 | 879 | [15][16][17] | 2017-05-19 |
Gene list
[edit]The following is a partial list of genes on human chromosome X. For complete list, see the link in the infobox on the right.
- AD16: encoding Alzheimer disease 16 protein
- AIC: encoding protein AIC
- APOO: encoding protein Apolipoprotein O
- ARMCX6: encoding protein Armadillo repeat containing X-linked 6
- BEX1: encoding protein Brain-expressed X-linked protein 1
- BEX2: encoding protein Brain-expressed X-linked protein 2
- BEX4: encoding protein Brain expressed, X-linked 4
- CCDC120: encoding protein Coiled coil domain containing protein 120
- CCDC22: encoding protein Coiled-coil domain containing 22
- CD99L2: CD99 antigen-like protein 2
- CDR1-AS: encoding protein CDR1 antisense RNA
- CFAP47: encoding protein Cilia and flagella associated protein 47
- CHRDL1: encoding protein Chordin-like 1
- CMTX2 encoding protein Charcot-Marie-Tooth neuropathy, X-linked 2 (recessive)
- CMTX3 encoding protein Charcot-Marie-Tooth neuropathy, X-linked 3 (dominant)
- CT45A5: encoding protein Cancer/testis antigen family 45, member A5
- CT55: encoding protein Cancer/testis antigen 55
- CXorf36: encoding protein hypothetical protein LOC79742
- CXorf57: encoding protein Chromosome X open reading frame 57
- CXorf40A: Chromosome X open reading frame 40
- CXorf49: chromosome X open reading frame 49. encoding protein
- CXorf66: encoding protein Chromosome X Open Reading Frame 66
- CXorf67: encoding protein Uncharacterized protein CXorf67
- DACH2: encoding protein Dachshund homolog 2
- EFHC2: encoding protein EF-hand domain (C-terminal) containing 2
- ERCC6L encoding protein ERCC excision repair 6 like, spindle assembly checkpoint helicase
- FAAH2: Fatty acid amide hydrolase 2
- F8A1: Factor VIII intron 22 protein
- FAM104B: encoding protein Family with sequence similarity 104 member B
- FAM120C: encoding protein Family with sequence similarity 120C
- FAM122B: Family with sequence similarity 122 member B
- FAM122C: encoding protein Family with sequence similarity 122C
- FAM127A: CAAX box protein 1
- FAM155B: encoding protein Family with Sequence Similarity 155 Member B
- FAM50A: Family with sequence similarity 50 member A
- FATE1: Fetal and adult testis-expressed transcript protein
- FMR1-AS1: encoding a long non-coding RNA FMR1 antisense RNA 1
- FRMPD3: encoding protein FERM and PDZ domain containing 3
- FRMPD4: encoding protein FERM and PDZ domain containing 4
- FUNDC1: encoding protein FUN14 domain containing 1
- FUNDC2: FUN14 domain-containing protein 2
- GAGE12F: encoding G antigen 12F protein
- GAGE2A: encoding G antigen 2A protein
- GATA1: encoding GATA1 transcription factor
- GNL3L encoding protein G protein nucleolar 3 like
- GPRASP2: G-protein coupled receptor-associated sorting protein 2
- GRIPAP1: encoding protein GRIP1-associated protein 1
- GRDX: encoding protein Graves disease, susceptibility to, X-linked
- HDHD1A: encoding enzyme Haloacid dehalogenase-like hydrolase domain-containing protein 1A
- HS6ST2: encoding protein Heparan sulfate 6-O-sulfotransferase 2
- ITM2A: encoding protein Integral membrane protein 2A
- LAS1L: encoding protein LAS1-like protein
- LOC101059915: encoding *LOC101059915 protein
- MAGEA2: encoding protein Melanoma-associated antigen 2
- MAGEA5: encoding protein Melanoma antigen family A, 5
- MAGEA8: encoding protein Melanoma antigen family A, 8
- MAGED4B: encoding protein Melanoma-associated antigen D4
- MAGT1: encoding protein Magnesium transporter protein 1
- MAGED4: encoding protein MAGE family member D4
- MAO-A: Monoamine Oxidase A
- MAO-B: Monoamine Oxidase B
- MAP3K15: encoding protein Mitogen-activated protein kinase kinase kinase 15
- MBNL3: encoding protein Muscleblind-like protein 3
- MBTPS2: encoding enzyme Membrane-bound transcription factor site-2 protease
- MCT-1: encoding protein MCTS1, re-initiation and release factor
- MIR106A: encoding microRNA MicroRNA 106
- MIR222: encoding microRNA MicroRNA 222
- MIR223: encoding protein MicroRNA 223
- MIR361: encoding microRNA MicroRNA 361
- MIR503: encoding microRNA MicroRNA 503
- MIR6087: encoding microRNA MicroRNA 6087
- MIR660: encoding microRNA MicroRNA 660
- MIRLET7F2: encoding protein MicroRNA let-7f-2
- MORF4L2: encoding protein Mortality factor 4-like protein 2
- MOSPD1: encoding protein Motile sperm domain containing 1
- MOSPD2: encoding protein Motile sperm domain containing 2
- NAP1L3: encoding protein Nucleosome assembly protein 1 like 3
- NBDY: encoding protein Negative regulator of P-body association
- NKRF: encoding protein NF-kappa-B-repressing factor
- NRK: encoding enzyme Nik-related protein kinase
- OPN1LW: long wave (red-cone) sensitive opsin
- OPN1MW: medium wave (green-cone) sensitive opsin
- OTUD5: encoding protein OTU deubiquitinase 5
- PASD1: encoding protein PAS domain-containing protein 1
- PAGE1: encoding protein PAGE family member 1
- PAGE2B: encoding PAGE family member 2B protein
- PBDC1: encoding a protein of unestablished function
- PCYT1B: encoding enzyme Choline-phosphate cytidylyltransferase B
- PIN4: encoding enzyme Peptidyl-prolyl cis-trans isomerase NIMA-interacting 4
- PLAC1: encoding protein Placenta-specific protein 1
- PLP2: encoding protein Proteolipid protein 2
- PRR32: encoding protein PRR32
- RPA4: encoding protein Replication protein A 30 kDa subunit
- RPS6KA6: encoding protein Ribosomal protein S6 kinase, 90kDa, polypeptide 6
- RRAGB: encoding protein Ras-related GTP-binding protein B
- RTL3: encoding protein Retrotransposon Gag like 3
- SFRS17A: encoding protein Splicing factor, arginine/serine-rich 17A
- SLC38A5: encoding protein Solute carrier family 38 member 5
- SLITRK2: encoding protein SLIT and NTRK-like protein 2
- SMARCA1: encoding protein Probable global transcription activator SNF2L1
- SMS: encoding enzyme Spermine synthase
- SPANXN1: encoding protein SPANX family member N1
- SPANXN5: encoding protein SPANX family member N5
- SPG16: encoding protein Spastic paraplegia 16 (complicated, X-linked recessive)
- SSR4: encoding protein Translocon-associated protein subunit delta
- TAF7L: encoding protein TATA-box binding protein associated factor 7-like
- TCEAL1: encoding protein Transcription elongation factor A protein-like 1
- TCEAL4: encoding protein Transcription elongation factor A protein-like 4
- TENT5D: encoding protein Terminal nucleotidyltransferase 5D
- TEX11: encoding protein Testis expressed 11
- THOC2: encoding protein THO complex subunit 2
- TMEM29: encoding protein Protein FAM156A
- TMEM47: encoding protein Transmembrane protein 47
- TMLHE: encoding enzyme Trimethyllysine dioxygenase, mitochondrial
- TNMD encoding protein Tenomodulin (also referred to as tendin, myodulin, Tnmd and TeM)
- TRAPPC2P1 encoding protein Trafficking protein particle complex subunit 2
- TREX2: encoding enzyme Three prime repair exonuclease 2
- TRO: encoding protein Trophinin
- TSPYL2: encoding protein Testis-specific Y-encoded-like protein 2
- TTC3P1: encoding protein Tetratricopeptide repeat domain 3 pseudogene 1
- USP51: encoding enzyme Ubiquitin carboxyl-terminal hydrolase 51
- VSIG1: encoding protein V-set and immunoglobulin domain containing 1
- YIPF6: encoding protein Protein YIPF6
- ZC3H12B: encoding protein ZC3H12B
- ZC4H2: encoding protein ZC4H2 Deficiency
- ZCCHC18: encoding protein Zinc finger CCHC-type containing 18
- ZFP92: encoding protein ZFP92 zinc finger protein
- ZMYM3: encoding protein Zinc finger MYM-type protein 3
- ZNF157: encoding protein Zinc finger protein 157
- ZNF182 encoding protein Zinc finger protein 182
- ZNF275: encoding protein Zinc finger protein 275
- ZNF674: encoding protein Zinc finger protein 674
Structure
[edit]It is theorized by Ross et al. 2005 and Ohno 1967 that the X chromosome is at least partially derived from the autosomal (non-sex-related) genome of other mammals, evidenced from interspecies genomic sequence alignments.
The X chromosome is notably larger and has a more active euchromatin region than its Y chromosome counterpart. Further comparison of the X and Y reveal regions of homology between the two. However, the corresponding region in the Y appears far shorter and lacks regions that are conserved in the X throughout primate species, implying a genetic degeneration for Y in that region. Because males have only one X chromosome, they are more likely to have an X chromosome-related disease.
It is estimated that about 10% of the genes encoded by the X chromosome are associated with a family of "CT" genes, so named because they encode for markers found in both tumor cells (in cancer patients) as well as in the human testis (in healthy patients).[18]
Role in disease
[edit]Numerical abnormalities
[edit]- Klinefelter syndrome is caused by the presence of one or more extra copies of the X chromosome in a male's cells.
- Males with Klinefelter syndrome typically have one extra copy of the X chromosome in each cell, for a total of two X chromosomes and one Y chromosome (47,XXY). It is less common for affected males to have two or three extra X chromosomes (48,XXXY or 49,XXXXY) or extra copies of both the X and Y chromosomes (48,XXYY) in each cell. The extra genetic material may lead to tall stature, learning and reading disabilities, and other medical problems. Each extra X chromosome lowers the child's IQ by about 15 points,[19][20] which means that the average IQ in Klinefelter syndrome is in general in the normal range, although below average. When additional X and/or Y chromosomes are present in 48,XXXY, 48,XXYY, or 49,XXXXY, developmental delays and cognitive difficulties can be more severe and mild intellectual disability may be present.
- Klinefelter syndrome can also result from an extra X chromosome in only some of the body's cells. These cases are called mosaic 46,XY/47,XXY.
- This syndrome results from an extra copy of the X chromosome in each of a female's cells. Females with trisomy X have three X chromosomes, for a total of 47 chromosomes per cell. The average IQ of females with this syndrome is 90, while the average IQ of unaffected siblings is 100.[21] Their stature on average is taller than normal females. They are fertile and their children do not inherit the condition.[22]
- Females with more than one extra copy of the X chromosome (48, tetrasomy X or 49, pentasomy X) have been identified, but these conditions are rare.
- This results when each of a female's cells has one normal X chromosome and the other sex chromosome is missing or altered. The missing genetic material affects development and causes the features of the condition, including short stature and infertility.
- About half of individuals with Turner syndrome have monosomy X (45,X), which means each cell in a woman's body has only one copy of the X chromosome instead of the usual two copies. Turner syndrome can also occur if one of the sex chromosomes is partially missing or rearranged rather than completely missing. Some women with Turner syndrome have a chromosomal change in only some of their cells. These cases are called Turner syndrome mosaics (45,X/46,XX).
X-linked recessive disorders
[edit]Sex linkage was first discovered in insects, e.g., T. H. Morgan's 1910 discovery of the pattern of inheritance of the white eyes mutation in Drosophila melanogaster.[23] Such discoveries helped to explain x-linked disorders in humans, e.g., haemophilia A and B, adrenoleukodystrophy, and red-green color blindness.
Other disorders
[edit]XX male syndrome is a rare disorder, where the SRY region of the Y chromosome has recombined to be located on one of the X chromosomes. As a result, the XX combination after fertilization has the same effect as a XY combination, resulting in a male. However, the other genes of the X chromosome cause feminization as well.
X-linked endothelial corneal dystrophy is an extremely rare disease of cornea associated with Xq25 region. Lisch epithelial corneal dystrophy is associated with Xp22.3.
Megalocornea 1 is associated with Xq21.3-q22[medical citation needed]
Adrenoleukodystrophy, a rare and fatal disorder that is carried by the mother on the x-cell. It affects only boys between the ages of 5 and 10 and destroys the protective cell surrounding the nerves, myelin, in the brain. The female carrier hardly shows any symptoms because females have a copy of the x-cell. This disorder causes a once healthy boy to lose all abilities to walk, talk, see, hear, and even swallow. Within 2 years after diagnosis, most boys with Adrenoleukodystrophy die.
Cytogenetic band
[edit]
| Chr. | Arm[28] | Band[29] | ISCN start[30] |
ISCN stop[30] |
Basepair start |
Basepair stop |
Stain[31] | Density |
|---|---|---|---|---|---|---|---|---|
| X | p | 22.33 | 0 | 323 | 1 | 4,400,000 | gneg | |
| X | p | 22.32 | 323 | 504 | 4,400,001 | 6,100,000 | gpos | 50 |
| X | p | 22.31 | 504 | 866 | 6,100,001 | 9,600,000 | gneg | |
| X | p | 22.2 | 866 | 1034 | 9,600,001 | 17,400,000 | gpos | 50 |
| X | p | 22.13 | 1034 | 1345 | 17,400,001 | 19,200,000 | gneg | |
| X | p | 22.12 | 1345 | 1448 | 19,200,001 | 21,900,000 | gpos | 50 |
| X | p | 22.11 | 1448 | 1577 | 21,900,001 | 24,900,000 | gneg | |
| X | p | 21.3 | 1577 | 1784 | 24,900,001 | 29,300,000 | gpos | 100 |
| X | p | 21.2 | 1784 | 1862 | 29,300,001 | 31,500,000 | gneg | |
| X | p | 21.1 | 1862 | 2120 | 31,500,001 | 37,800,000 | gpos | 100 |
| X | p | 11.4 | 2120 | 2430 | 37,800,001 | 42,500,000 | gneg | |
| X | p | 11.3 | 2430 | 2624 | 42,500,001 | 47,600,000 | gpos | 75 |
| X | p | 11.23 | 2624 | 2948 | 47,600,001 | 50,100,000 | gneg | |
| X | p | 11.22 | 2948 | 3129 | 50,100,001 | 54,800,000 | gpos | 25 |
| X | p | 11.21 | 3129 | 3206 | 54,800,001 | 58,100,000 | gneg | |
| X | p | 11.1 | 3206 | 3297 | 58,100,001 | 61,000,000 | acen | |
| X | q | 11.1 | 3297 | 3491 | 61,000,001 | 63,800,000 | acen | |
| X | q | 11.2 | 3491 | 3620 | 63,800,001 | 65,400,000 | gneg | |
| X | q | 12 | 3620 | 3827 | 65,400,001 | 68,500,000 | gpos | 50 |
| X | q | 13.1 | 3827 | 4137 | 68,500,001 | 73,000,000 | gneg | |
| X | q | 13.2 | 4137 | 4292 | 73,000,001 | 74,700,000 | gpos | 50 |
| X | q | 13.3 | 4292 | 4447 | 74,700,001 | 76,800,000 | gneg | |
| X | q | 21.1 | 4447 | 4732 | 76,800,001 | 85,400,000 | gpos | 100 |
| X | q | 21.2 | 4732 | 4809 | 85,400,001 | 87,000,000 | gneg | |
| X | q | 21.31 | 4809 | 5107 | 87,000,001 | 92,700,000 | gpos | 100 |
| X | q | 21.32 | 5107 | 5184 | 92,700,001 | 94,300,000 | gneg | |
| X | q | 21.33 | 5184 | 5430 | 94,300,001 | 99,100,000 | gpos | 75 |
| X | q | 22.1 | 5430 | 5701 | 99,100,001 | 103,300,000 | gneg | |
| X | q | 22.2 | 5701 | 5843 | 103,300,001 | 104,500,000 | gpos | 50 |
| X | q | 22.3 | 5843 | 6050 | 104,500,001 | 109,400,000 | gneg | |
| X | q | 23 | 6050 | 6322 | 109,400,001 | 117,400,000 | gpos | 75 |
| X | q | 24 | 6322 | 6619 | 117,400,001 | 121,800,000 | gneg | |
| X | q | 25 | 6619 | 7059 | 121,800,001 | 129,500,000 | gpos | 100 |
| X | q | 26.1 | 7059 | 7253 | 129,500,001 | 131,300,000 | gneg | |
| X | q | 26.2 | 7253 | 7395 | 131,300,001 | 134,500,000 | gpos | 25 |
| X | q | 26.3 | 7395 | 7602 | 134,500,001 | 138,900,000 | gneg | |
| X | q | 27.1 | 7602 | 7808 | 138,900,001 | 141,200,000 | gpos | 75 |
| X | q | 27.2 | 7808 | 7886 | 141,200,001 | 143,000,000 | gneg | |
| X | q | 27.3 | 7886 | 8145 | 143,000,001 | 148,000,000 | gpos | 100 |
| X | q | 28 | 8145 | 8610 | 148,000,001 | 156,040,895 | gneg |
Research
[edit]In July 2020 scientists reported the first complete and gap-less assembly of a human X chromosome.[32][33]
See also
[edit]References
[edit]- Earlier versions of this article contain material from the National Library of Medicine, a part of the National Institutes of Health (USA), which, as a US government publication, is in the public domain.
- ^ Tom Strachan; Andrew Read (2 April 2010). Human Molecular Genetics. Garland Science. p. 45. ISBN 978-1-136-84407-2.
- ^ a b c Genome Decoration Page, NCBI. Ideogram data for Homo sapience (850 bphs, Assembly GRCh38.p3). Last update 2014-06-03. Retrieved 2017-04-26.
- ^ Angier, Natalie (2007-05-01). "For Motherly X Chromosome, Gender Is Only the Beginning". The New York Times. Retrieved 2007-05-01.
- ^ a b James Schwartz, In Pursuit of the Gene: From Darwin to DNA, pages 155-158, Harvard University Press, 2009 ISBN 0674034910
- ^ David Bainbridge, 'The X in Sex: How the X Chromosome Controls Our Lives, pages 3-5, Harvard University Press, 2003 ISBN 0674016211.
- ^ Bainbridge, pages 65-66
- ^ a b Hutchison, Luke (September 2004). "Growing the Family Tree: The Power of DNA in Reconstructing Family Relationships" (PDF). Proceedings of the First Symposium on Bioinformatics and Biotechnology (BIOT-04). Retrieved 2016-09-03.
- ^ Carrel L, Willard H (2005). "X-inactivation profile reveals extensive variability in X-linked gene expression in females". Nature. 434 (7031): 400–4. Bibcode:2005Natur.434..400C. doi:10.1038/nature03479. PMID 15772666. S2CID 4358447.
- ^ Veneti Z, Gkouskou KK, Eliopoulos AG (July 2017). "Polycomb Repressor Complex 2 in Genomic Instability and Cancer". Int J Mol Sci. 18 (8): 1657. doi:10.3390/ijms18081657. PMC 5578047. PMID 28758948.
- ^ Pertea M, Salzberg SL (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biol. 11 (5): 206. doi:10.1186/gb-2010-11-5-206. PMC 2898077. PMID 20441615.
- ^ "Search results - X[CHR] AND "Homo sapiens"[Organism] AND ("has ccds"[Properties] AND alive[prop]) - Gene". NCBI. CCDS Release 20 for Homo sapiens. 2016-09-08. Retrieved 2017-05-28.
- ^ "Statistics & Downloads for chromosome X". HUGO Gene Nomenclature Committee. 2017-05-12. Archived from the original on 2017-06-29. Retrieved 2017-05-19.
- ^ "Chromosome X: Chromosome summary - Homo sapiens". Ensembl Release 88. 2017-03-29. Retrieved 2017-05-19.
- ^ "Human chromosome X: entries, gene names and cross-references to MIM". UniProt. 2018-02-28. Retrieved 2018-03-16.
- ^ "Search results - X[CHR] AND "Homo sapiens"[Organism] AND ("genetype protein coding"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ "Search results - X[CHR] AND "Homo sapiens"[Organism] AND ( ("genetype miscrna"[Properties] OR "genetype ncrna"[Properties] OR "genetype rrna"[Properties] OR "genetype trna"[Properties] OR "genetype scrna"[Properties] OR "genetype snrna"[Properties] OR "genetype snorna"[Properties]) NOT "genetype protein coding"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ "Search results - X[CHR] AND "Homo sapiens"[Organism] AND ("genetype pseudo"[Properties] AND alive[prop]) - Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ Ross M, et al. (2005). "The DNA sequence of the human X chromosome". Nature. 434 (7031): 325–37. Bibcode:2005Natur.434..325R. doi:10.1038/nature03440. PMC 2665286. PMID 15772651.
- ^ Harold Chen; Ian Krantz; Mary L Windle; Margaret M McGovern; Paul D Petry; Bruce Buehler (2013-02-22). "Klinefelter Syndrome Pathophysiology". Medscape. Retrieved 2014-07-18.
- ^ Visootsak J, Graham JM (2006). "Klinefelter syndrome and other sex chromosomal aneuploidies". Orphanet J Rare Dis. 1: 42. doi:10.1186/1750-1172-1-42. PMC 1634840. PMID 17062147.
- ^ Bender B, Puck M, Salbenblatt J, Robinson A (1986). Smith S (ed.). Cognitive development of children with sex chromosome abnormalities. San Diego: College Hill Press. pp. 175–201.
- ^ "Triple X syndrome". Genetics Home Reference. 2014-07-14. Retrieved 2014-07-18.
- ^ Morgan, T. H. (1910). "Sex-limited inheritance in Drosophila". Science. 32 (812): 120–122. Bibcode:1910Sci....32..120M. doi:10.1126/science.32.812.120. PMID 17759620.
- ^ Genome Decoration Page, NCBI. Ideogram data for Homo sapience (400 bphs, Assembly GRCh38.p3). Last update 2014-03-04. Retrieved 2017-04-26.
- ^ Genome Decoration Page, NCBI. Ideogram data for Homo sapience (550 bphs, Assembly GRCh38.p3). Last update 2015-08-11. Retrieved 2017-04-26.
- ^ International Standing Committee on Human Cytogenetic Nomenclature (2013). ISCN 2013: An International System for Human Cytogenetic Nomenclature (2013). Karger Medical and Scientific Publishers. ISBN 978-3-318-02253-7.
- ^ Sethakulvichai, W.; Manitpornsut, S.; Wiboonrat, M.; Lilakiatsakun, W.; Assawamakin, A.; Tongsima, S. (2012). "Estimation of band level resolutions of human chromosome images". 2012 Ninth International Conference on Computer Science and Software Engineering (JCSSE). pp. 276–282. doi:10.1109/JCSSE.2012.6261965. ISBN 978-1-4673-1921-8. S2CID 16666470.
- ^ "p": Short arm; "q": Long arm.
- ^ For cytogenetic banding nomenclature, see article locus.
- ^ a b These values (ISCN start/stop) are based on the length of bands/ideograms from the ISCN book, An International System for Human Cytogenetic Nomenclature (2013). Arbitrary unit.
- ^ gpos: Region which is positively stained by G banding, generally AT-rich and gene poor; gneg: Region which is negatively stained by G banding, generally CG-rich and gene rich; acen Centromere. var: Variable region; stalk: Stalk.
- ^ "Scientists achieve first complete assembly of human X chromosome". phys.org. Retrieved 16 August 2020.
- ^ Miga, Karen H.; Koren, Sergey; Rhie, Arang; Vollger, Mitchell R.; Gershman, Ariel; Bzikadze, Andrey; Brooks, Shelise; Howe, Edmund; Porubsky, David; Logsdon, Glennis A.; Schneider, Valerie A.; Potapova, Tamara; Wood, Jonathan; Chow, William; Armstrong, Joel; Fredrickson, Jeanne; Pak, Evgenia; Tigyi, Kristof; Kremitzki, Milinn; Markovic, Christopher; Maduro, Valerie; Dutra, Amalia; Bouffard, Gerard G.; Chang, Alexander M.; Hansen, Nancy F.; Wilfert, Amy B.; Thibaud-Nissen, Françoise; Schmitt, Anthony D.; Belton, Jon-Matthew; Selvaraj, Siddarth; Dennis, Megan Y.; Soto, Daniela C.; Sahasrabudhe, Ruta; Kaya, Gulhan; Quick, Josh; Loman, Nicholas J.; Holmes, Nadine; Loose, Matthew; Surti, Urvashi; Risques, Rosa ana; Lindsay, Tina A. Graves; Fulton, Robert; Hall, Ira; Paten, Benedict; Howe, Kerstin; Timp, Winston; Young, Alice; Mullikin, James C.; Pevzner, Pavel A.; Gerton, Jennifer L.; Sullivan, Beth A.; Eichler, Evan E.; Phillippy, Adam M. (14 July 2020). "Telomere-to-telomere assembly of a complete human X chromosome". Nature. 585 (7823): 79–84. Bibcode:2020Natur.585...79M. doi:10.1038/s41586-020-2547-7. ISSN 1476-4687. PMC 7484160. PMID 32663838.
External links
[edit]- National Institutes of Health. "X chromosome". Genetics Home Reference. Archived from the original on 2007-07-08. Retrieved 2017-05-06.
- "X chromosome". Human Genome Project Information Archive 1990–2003. Retrieved 2017-05-06.






