User:TessaColameta/sandbox
 | This is a user sandbox of TessaColameta. You can use it for testing or practicing edits. This is not the sandbox where you should draft your assigned article for a dashboard.wikiedu.org course. To find the right sandbox for your assignment, visit your Dashboard course page and follow the Sandbox Draft link for your assigned article in the My Articles section. |
Group Draft Space: User:GomezChristian/sandbox
| chorismate mutase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Reaction catalyzed by chorismate mutase | |||||||||
| Identifiers | |||||||||
| EC no. | 5.4.99.5 | ||||||||
| CAS no. | 9068-30-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
In enzymology, a chorismate mutase (EC 5.4.99.5) is an enzyme that catalyzes the chemical reaction for the conversion of chorismate to prephenate in the pathway to the production of phenylalanine and tyrosine, also known as the shikimate pathway. Hence, this enzyme has one substrate, chorismate, and one product, prephenate. Chorismate mutase is found at a branch point in the pathway. The enzyme channels the substrate, chorismate to the biosynthesis of tyrosine and phenylalanine and away from tryptophan.[1] Its role in maintaining the balance of these aromatic amino acids in the cell is vital.[2] This is the single known example of a naturally occurring enzyme catalyzing a pericyclic reaction.[2][nb 1] Chorismate mutase(CM) is only found in fungi, bacteria, and higher plants. This protein may use the morpheein model of allosteric regulation.[4]
Protein family
[edit]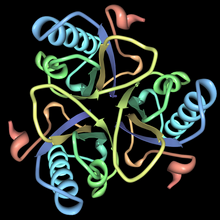
This enzyme belongs to the family of isomerases, specifically those intramolecular transferases transferring other groups. The systematic name of this enzyme class is chorismate pyruvatemutase. Chorismate mutase, also known as hydroxyphenylpyruvate synthase, participates in phenylalanine, tyrosine and tryptophan biosynthesis[1]. The structures of chorismate mutases vary in different organisms, but the majority belong to the AroQ family and are characterized by an intertwined homodimer of 3-helical subunits. Most chorismate mutases in this family look similar to that of Escherichia coli. For example, the secondary structure of the chorismate mutase of yeast is very similar to that of E. coli. Chorimate mutase in the AroQ family are more common in nature and are widely distributed among the prokaryotes[1]. For optimal function, they usually have to be accompanied by another enzyme such as prephanate dehydrogenase. These chorismate mutases are typically bifunctional enzymes, meaning they contain two catalytic capacities in the same polypeptide chain[1]. However, the chorismate mutase of eukaryotic organisms are more commonly monofunctional. There are organisms such as Bacillus subtilis whose chorismate mutase have a completely different structure and are monofunctional. These enzymes belong to the AroH family and are characterized by a trimeric α/β barrel topology.
Mechanism of catalysis
[edit]The mechanism for the transformation of chorismate to prephenate is formally a Claisen rearrangement. This transformation is the first committed step in the pathway to production of the aromatic amino acids: tyrosine and phenylalanine. In the absence of enzyme catalysis this mechanism proceeds as a concerted, but asynchronous step and is an exergonic process.[2] As a result, there is no formal intermediate, but rather a chair-like transition state. The addition of chorismate mutase, increases the rate of the reaction a million fold. There have been extensive studies on the exact mechanism of this reaction, but the rate-determining step has yet to be uncovered. Some questions that remain surrounding the mechanism are how conformational constraint of the flexible substrate, specific hydrogen bonding to the transition state, and electrostatic interactions actually contribute to catalysis. One study using CM from B. subtilis showed evidence that when a cation was aptly placed in the active site, the electrostatic interactions between it and the negatively charged transition state promoted catalysis.[2]
Notes
[edit]- ^ Dimethylallyltryptophan synthase has been proposed to catalyze a Cope rearrangement, but this has yet to be proven definitively[3]
References
[edit]- ^ a b c d Qamra R, Prakash P, Aruna B, Hasnain SE, Mande SC (June 2006). "The 2.15 A crystal structure of Mycobacterium tuberculosis chorismate mutase reveals an unexpected gene duplication and suggests a role in host-pathogen interactions". Biochemistry. 45 (23): 6997–7005. doi:10.1021/bi0606445. PMID 16752890.
- ^ a b c d Kast P, Grisostomi C, Chen IA, Li S, Krengel U, Xue Y, Hilvert D (November 2000). "A strategically positioned cation is crucial for efficient catalysis by chorismate mutase". The Journal of Biological Chemistry. 275 (47): 36832–8. doi:10.1074/jbc.M006351200. PMID 10960481.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Luk LY, Qian Q, Tanner ME (August 2011). "A cope rearrangement in the reaction catalyzed by dimethylallyltryptophan synthase?". Journal of the American Chemical Society. 133 (32): 12342–5. doi:10.1021/ja2034969. PMID 21766851.
- ^ Selwood T, Jaffe EK (March 2012). "Dynamic dissociating homo-oligomers and the control of protein function". Archives of Biochemistry and Biophysics. 519 (2): 131–43. doi:10.1016/j.abb.2011.11.020. PMC 3298769. PMID 22182754.
{{cite journal}}: Unknown parameter|last-author-amp=ignored (|name-list-style=suggested) (help)
