Bovine pancreatic ribonuclease
| Pancreatic ribonuclease | |||||||||
|---|---|---|---|---|---|---|---|---|---|
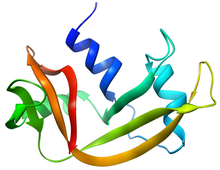 Structure of RNase A | |||||||||
| Identifiers | |||||||||
| EC no. | 3.1.27.5 | ||||||||
| CAS no. | 9001-99-4 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||

Bovine pancreatic ribonuclease, also often referred to as bovine pancreatic ribonuclease A or simply RNase A, is a pancreatic ribonuclease enzyme that cleaves single-stranded RNA. Bovine pancreatic ribonuclease is one of the classic model systems of protein science.[1] Two Nobel Prizes in Chemistry have been awarded in recognition of work on bovine pancreatic ribonuclease: in 1972, the Prize was awarded to Christian Anfinsen for his work on protein folding and to Stanford Moore and William Stein for their work on the relationship between the protein's structure and its chemical mechanism;[2] in 1984, the Prize was awarded to Robert Bruce Merrifield for development of chemical synthesis of proteins.[3]
History
[edit]Bovine pancreatic ribonuclease became a common model system in the study of proteins largely because it was extremely stable and could be purified in large quantities. In the 1940s Armour and Company purified a kilogram of protein - a very large quantity, particularly by the protein purification standards of the time - and offered samples at low cost to interested scientists.[4] The ability to have a single lot of purified enzyme made it a predominant model system for protein studies. It remains commonly referred to as ribonuclease A or RNase A as the most prominent member of its protein family, known variously as pancreatic ribonuclease, ribonuclease A, or ribonuclease I.
Christian Anfinsen's studies of the oxidative folding process of bovine pancreatic ribonuclease laid the groundwork for understanding the relationship between amino acid sequence and a protein's folded three-dimensional structure and solidified the thermodynamic hypothesis of protein folding, according to which the folded form of a protein represents its free energy minimum.[4][5]
RNase A was the first enzyme for which a correct catalytic mechanism was proposed, even before its structure was known.[6] RNase A was the first protein for showing the effects of non-native isomers of peptide bonds preceding proline residues in protein folding.[7]
Bovine pancreatic ribonuclease was also the model protein used to work out many spectroscopic methods for assaying protein structure, including absorbance, circular dichroism, Raman, electron paramagnetic resonance (EPR) and nuclear magnetic resonance (NMR) spectroscopy. It was the first model protein for the development of chemical methods for the study of proteins, such as chemical modification of exposed side chains, antigenic recognition, and limited proteolysis of disordered segments. Ribonuclease S, which is RNase A that has been treated with the protease subtilisin, was the third protein to have its crystallographic structure solved, in 1967.[8]
Structure and properties
[edit]
RNase A is a relatively small protein (124 residues, ~13.7 kDa). It can be characterized as a two-layer protein with a deep cleft for binding the RNA substrate. The first layer is composed of three alpha helices (residues 3-13, 24-34 and 50-60) from the N-terminal half of the protein. The second layer consist of three β-hairpins (residues 61-74, 79-104 and 105-124 from the C-terminal half) arranged in two β-sheets. The hairpins 61-74 and 105-124 form a four-stranded, antiparallel β-sheet that lies on helix 3 (residues 50-60). The longest β-hairpin 79-104 mates with a short β-strand (residues 42-45) to form a three-stranded, antiparallel β-sheet that lies on helix 2 (residues 24-34).
RNase A has four disulfide bonds in its native state: Cys26-Cys84, Cys58-110, Cys40-95 and Cys65-72. The first two (26-84 and 58-110) are essential for conformational folding; each joins an alpha helix of the first layer to a beta sheet of the second layer, forming a small hydrophobic core in its vicinity. The latter two disulfide bonds (40-95 and 65-72) are less essential for folding; either one can be reduced (but not both) without affecting the native structure under physiological conditions. These disulfide bonds connect loop segments and are relatively exposed to solvent. The 65-72 disulfide bond has an extraordinarily high propensity to form, significantly more than would be expected from its loop entropy, both as a peptide and in the full-length protein. This suggests that the 61-74 β-hairpin has a high propensity to fold conformationally.
RNase A is a basic protein (pI = 9.63); its many positive charges are consistent with its binding to RNA (a poly-anion). More generally, RNase A is unusually polar or, rather, unusually lacking in hydrophobic groups, especially aliphatic ones. This may account for its need of four disulfide bonds to stabilize its structure. The low hydrophobic content may also serve to reduce the physical repulsion between highly charged groups (its own and those of its substrate RNA) and regions of low dielectric constant (the nonpolar residues).
The N-terminal α-helix of RNase A (residues 3-13) is connected to the rest of RNase A by a flexible linker (residues 16-23). As shown by F. M. Richards, this linker may be cleaved by subtilisin between residues 20 and 21 without causing the N-terminal helix to dissociate from the rest of RNase A. The peptide-protein complex is called "RNase S", the peptide (residues 1-20) is called the "S-peptide" and the remainder (residues 21-124) is called the "S-protein". The dissociation constant of the S-peptide for the S-protein is roughly 30 pM; this tight binding can be exploited for protein purification by attaching the S-peptide to the protein of interest and passing a mixture over an affinity column with bound S-protein. [A smaller C-peptide (residues 1-13) also works.] The RNase S model system has also been used for studying protein folding by coupling folding and association. The S-peptide was the first peptide from a native protein shown to have (flickering) secondary structure in isolation (by Klee and Brown in 1967).
RNase A cleaves specifically after pyrimidine nucleotides.[9] Cleavage takes place in two steps: first, the 3’,5’-phosphodiester bond is cleaved to generate a 2’,3’-cyclic phosphodiester intermediate; second, the cyclic phosphodiester is hydrolyzed to a 3’-monophosphate.[10] It can be inhibited by ribonuclease inhibitor protein, by heavy metal ions, and by uridine-vanadate complexes.[10]
Enzymatic mechanism
[edit]The positive charges of RNase A lie mainly in a deep cleft between two lobes. The RNA substrate lies in this cleft and is cleaved by two catalytic histidine residues, His12 and His119, to form a 2',3'-cyclic phosphate intermediate that is stabilized by nearby Lys41.
Enzyme regulation
[edit]This protein may use the morpheein model of allosteric regulation.[11]
See also
[edit]References
[edit]- ^ Raines RT (1998). "Ribonuclease A". Chem. Rev. 98 (3): 1045–1066. doi:10.1021/cr960427h. PMID 11848924.
- ^ "The Nobel Prize in Chemistry 1972". Nobelprize.org. Retrieved 10 February 2015.
- ^ "The Nobel Prize in Chemistry 1984". Nobelprize.org. Retrieved 10 February 2015.
- ^ a b Richards FM (1972). "The 1972 nobel prize for chemistry". Science. 178 (4060): 492–3. Bibcode:1972Sci...178..492R. doi:10.1126/science.178.4060.492. PMID 17754377.
- ^ Marshall, G. R.; Feng, J. A.; Kuster, D. J. (2008). "Back to the future: Ribonuclease A". Biopolymers. 90 (3): 259–77. doi:10.1002/bip.20845. PMID 17868092.
- ^ Cuchillo CM, Nogués MV, Raines RT (2011). "Bovine pancreatic ribonuclease: fifty years of the first enzymatic reaction mechanism". Biochemistry. 50 (37): 7835–7841. doi:10.1021/bi201075b. PMC 3172371. PMID 21838247.
- ^ Schmid, FX; Baldwin, RL (October 1978). "Acid catalysis of the formation of the slow-folding species of RNase A: evidence that the reaction is proline isomerization". Proceedings of the National Academy of Sciences of the United States of America. 75 (10): 4764–8. Bibcode:1978PNAS...75.4764S. doi:10.1073/pnas.75.10.4764. PMC 336200. PMID 283390.
- ^ Wyckoff HW, Hardman KD, Allewell NM, Inagami T, Johnson LN, Richards FM (1967). "The structure of ribonuclease-S at 3.5 A resolution". J. Biol. Chem. 242 (17): 3984–8. doi:10.1016/S0021-9258(18)95844-8. PMID 6037556.
- ^ Volkin E, Cohn WE (1953). "On the structure of ribonucleic acids. II. The products of ribonuclease action". J. Biol. Chem. 205 (2): 767–82. doi:10.1016/S0021-9258(18)49221-6. PMID 13129256.
- ^ a b Krystal Worthington. "Ribonuclease - Worthington Enzyme Manual". Retrieved 2011-09-26.
- ^ Selwood T, Jaffe EK (2012). "Dynamic dissociating homo-oligomers and the control of protein function". Arch. Biochem. Biophys. 519 (2): 131–43. doi:10.1016/j.abb.2011.11.020. PMC 3298769. PMID 22182754.
Further reading
[edit]- Kartha, G.; Bello, J.; Harker, D. (1967). Tertiary Structure of Ribonuclease. Boston: Nature. ISBN 0-12-588945-3.
- Scheraga HA, Wedemeyer WJ, Welker E (2001). "Bovine Pancreatic Ribonuclease A: Oxidative and Conformational Folding Studies". Ribonucleases - Part A. Methods in Enzymology. Vol. 341. pp. 189–221. doi:10.1016/S0076-6879(01)41153-0. ISBN 9780121822422. PMID 11582778.
External links
[edit]- Ribonuclease+A at the U.S. National Library of Medicine Medical Subject Headings (MeSH)

