Biomolecule
| Part of a series on |
| Biochemistry |
|---|
 |

A biomolecule or biological molecule is loosely defined as a molecule produced by a living organism and essential to one or more typically biological processes.[1] Biomolecules include large macromolecules such as proteins, carbohydrates, lipids, and nucleic acids, as well as small molecules (Micromolecules) such as vitamins and hormones. A general name for this class of material is biological materials. Biomolecules are an important element of living organisms, those biomolecules are often endogenous,[2] produced within the organism[3] but organisms usually need exogenous biomolecules, for example certain nutrients, to survive.
Biology and its subfields of biochemistry and molecular biology study biomolecules and their reactions. Most biomolecules are organic compounds, and just four elements—oxygen, carbon, hydrogen, and nitrogen—make up 96% of the human body's mass. But many other elements, such as the various biometals, are also present in small amounts.
The uniformity of both specific types of molecules (the biomolecules) and of certain metabolic pathways are invariant features among the wide diversity of life forms; thus these biomolecules and metabolic pathways are referred to as "biochemical universals"[4] or "theory of material unity of the living beings", a unifying concept in biology, along with cell theory and evolution theory.[5]
Types of biomolecules
[edit]A diverse range of biomolecules exist, including:
- Small molecules:
- Monomers, oligomers and polymers:
| Biomonomers | Bio-oligo | Biopolymers | Polymerization process | Covalent bond name between monomers |
|---|---|---|---|---|
| Amino acids | Oligopeptides | Polypeptides, proteins (hemoglobin...) | Polycondensation | Peptide bond |
| Monosaccharides | Oligosaccharides | Polysaccharides (cellulose...) | Polycondensation | Glycosidic bond |
| Isoprene | Terpenes | Polyterpenes: cis-1,4-polyisoprene natural rubber and trans-1,4-polyisoprene gutta-percha | Polyaddition |
Nucleosides and nucleotides
[edit]Nucleosides are molecules formed by attaching a nucleobase to a ribose or deoxyribose ring. Examples of these include cytidine (C), uridine (U), adenosine (A), guanosine (G), and thymidine (T).
Nucleosides can be phosphorylated by specific kinases in the cell, producing nucleotides. Both DNA and RNA are polymers, consisting of long, linear molecules assembled by polymerase enzymes from repeating structural units, or monomers, of mononucleotides. DNA uses the deoxynucleotides C, G, A, and T, while RNA uses the ribonucleotides (which have an extra hydroxyl(OH) group on the pentose ring) C, G, A, and U. Modified bases are fairly common (such as with methyl groups on the base ring), as found in ribosomal RNA or transfer RNAs or for discriminating the new from old strands of DNA after replication.[6]
Each nucleotide is made of an acyclic nitrogenous base, a pentose and one to three phosphate groups. They contain carbon, nitrogen, oxygen, hydrogen and phosphorus. They serve as sources of chemical energy (adenosine triphosphate and guanosine triphosphate), participate in cellular signaling (cyclic guanosine monophosphate and cyclic adenosine monophosphate), and are incorporated into important cofactors of enzymatic reactions (coenzyme A, flavin adenine dinucleotide, flavin mononucleotide, and nicotinamide adenine dinucleotide phosphate).[7]
DNA and RNA structure
[edit]DNA structure is dominated by the well-known double helix formed by Watson-Crick base-pairing of C with G and A with T. This is known as B-form DNA, and is overwhelmingly the most favorable and common state of DNA; its highly specific and stable base-pairing is the basis of reliable genetic information storage. DNA can sometimes occur as single strands (often needing to be stabilized by single-strand binding proteins) or as A-form or Z-form helices, and occasionally in more complex 3D structures such as the crossover at Holliday junctions during DNA replication.[7]
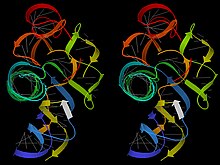
RNA, in contrast, forms large and complex 3D tertiary structures reminiscent of proteins, as well as the loose single strands with locally folded regions that constitute messenger RNA molecules. Those RNA structures contain many stretches of A-form double helix, connected into definite 3D arrangements by single-stranded loops, bulges, and junctions.[8] Examples are tRNA, ribosomes, ribozymes, and riboswitches. These complex structures are facilitated by the fact that RNA backbone has less local flexibility than DNA but a large set of distinct conformations, apparently because of both positive and negative interactions of the extra OH on the ribose.[9] Structured RNA molecules can do highly specific binding of other molecules and can themselves be recognized specifically; in addition, they can perform enzymatic catalysis (when they are known as "ribozymes", as initially discovered by Tom Cech and colleagues).[10]
Saccharides
[edit]Monosaccharides are the simplest form of carbohydrates with only one simple sugar. They essentially contain an aldehyde or ketone group in their structure.[11] The presence of an aldehyde group in a monosaccharide is indicated by the prefix aldo-. Similarly, a ketone group is denoted by the prefix keto-.[6] Examples of monosaccharides are the hexoses, glucose, fructose, Trioses, Tetroses, Heptoses, galactose, pentoses, ribose, and deoxyribose. Consumed fructose and glucose have different rates of gastric emptying, are differentially absorbed and have different metabolic fates, providing multiple opportunities for two different saccharides to differentially affect food intake.[11] Most saccharides eventually provide fuel for cellular respiration.
Disaccharides are formed when two monosaccharides, or two single simple sugars, form a bond with removal of water. They can be hydrolyzed to yield their saccharin building blocks by boiling with dilute acid or reacting them with appropriate enzymes.[6] Examples of disaccharides include sucrose, maltose, and lactose.
Polysaccharides are polymerized monosaccharides, or complex carbohydrates. They have multiple simple sugars. Examples are starch, cellulose, and glycogen. They are generally large and often have a complex branched connectivity. Because of their size, polysaccharides are not water-soluble, but their many hydroxy groups become hydrated individually when exposed to water, and some polysaccharides form thick colloidal dispersions when heated in water.[6] Shorter polysaccharides, with 3 to 10 monomers, are called oligosaccharides.[12] A fluorescent indicator-displacement molecular imprinting sensor was developed for discriminating saccharides. It successfully discriminated three brands of orange juice beverage.[13] The change in fluorescence intensity of the sensing films resulting is directly related to the saccharide concentration.[14]
Lignin
[edit]Lignin is a complex polyphenolic macromolecule composed mainly of beta-O4-aryl linkages. After cellulose, lignin is the second most abundant biopolymer and is one of the primary structural components of most plants. It contains subunits derived from p-coumaryl alcohol, coniferyl alcohol, and sinapyl alcohol,[15] and is unusual among biomolecules in that it is racemic. The lack of optical activity is due to the polymerization of lignin which occurs via free radical coupling reactions in which there is no preference for either configuration at a chiral center.
Lipid
[edit]Lipids (oleaginous) are chiefly fatty acid esters, and are the basic building blocks of biological membranes. Another biological role is energy storage (e.g., triglycerides). Most lipids consist of a polar or hydrophilic head (typically glycerol) and one to three non polar or hydrophobic fatty acid tails, and therefore they are amphiphilic. Fatty acids consist of unbranched chains of carbon atoms that are connected by single bonds alone (saturated fatty acids) or by both single and double bonds (unsaturated fatty acids). The chains are usually 14-24 carbon groups long, but it is always an even number.
For lipids present in biological membranes, the hydrophilic head is from one of three classes:
- Glycolipids, whose heads contain an oligosaccharide with 1-15 saccharide residues.
- Phospholipids, whose heads contain a positively charged group that is linked to the tail by a negatively charged phosphate group.
- Sterols, whose heads contain a planar steroid ring, for example, cholesterol.
Other lipids include prostaglandins and leukotrienes which are both 20-carbon fatty acyl units synthesized from arachidonic acid. They are also known as fatty acids
Amino acids
[edit]Amino acids contain both amino and carboxylic acid functional groups. (In biochemistry, the term amino acid is used when referring to those amino acids in which the amino and carboxylate functionalities are attached to the same carbon, plus proline which is not actually an amino acid).
Modified amino acids are sometimes observed in proteins; this is usually the result of enzymatic modification after translation (protein synthesis). For example, phosphorylation of serine by kinases and dephosphorylation by phosphatases is an important control mechanism in the cell cycle. Only two amino acids other than the standard twenty are known to be incorporated into proteins during translation, in certain organisms:
- Selenocysteine is incorporated into some proteins at a UGA codon, which is normally a stop codon.
- Pyrrolysine is incorporated into some proteins at a UAG codon. For instance, in some methanogens in enzymes that are used to produce methane.
Besides those used in protein synthesis, other biologically important amino acids include carnitine (used in lipid transport within a cell), ornithine, GABA and taurine.
Protein structure
[edit]The particular series of amino acids that form a protein is known as that protein's primary structure. This sequence is determined by the genetic makeup of the individual. It specifies the order of side-chain groups along the linear polypeptide "backbone".
Proteins have two types of well-classified, frequently occurring elements of local structure defined by a particular pattern of hydrogen bonds along the backbone: alpha helix and beta sheet. Their number and arrangement is called the secondary structure of the protein. Alpha helices are regular spirals stabilized by hydrogen bonds between the backbone CO group (carbonyl) of one amino acid residue and the backbone NH group (amide) of the i+4 residue. The spiral has about 3.6 amino acids per turn, and the amino acid side chains stick out from the cylinder of the helix. Beta pleated sheets are formed by backbone hydrogen bonds between individual beta strands each of which is in an "extended", or fully stretched-out, conformation. The strands may lie parallel or antiparallel to each other, and the side-chain direction alternates above and below the sheet. Hemoglobin contains only helices, natural silk is formed of beta pleated sheets, and many enzymes have a pattern of alternating helices and beta-strands. The secondary-structure elements are connected by "loop" or "coil" regions of non-repetitive conformation, which are sometimes quite mobile or disordered but usually adopt a well-defined, stable arrangement.[16]
The overall, compact, 3D structure of a protein is termed its tertiary structure or its "fold". It is formed as result of various attractive forces like hydrogen bonding, disulfide bridges, hydrophobic interactions, hydrophilic interactions, van der Waals force etc.
When two or more polypeptide chains (either of identical or of different sequence) cluster to form a protein, quaternary structure of protein is formed. Quaternary structure is an attribute of polymeric (same-sequence chains) or heteromeric (different-sequence chains) proteins like hemoglobin, which consists of two "alpha" and two "beta" polypeptide chains.
Apoenzymes
[edit]An apoenzyme (or, generally, an apoprotein) is the protein without any small-molecule cofactors, substrates, or inhibitors bound. It is often important as an inactive storage, transport, or secretory form of a protein. This is required, for instance, to protect the secretory cell from the activity of that protein. Apoenzymes become active enzymes on addition of a cofactor. Cofactors can be either inorganic (e.g., metal ions and iron-sulfur clusters) or organic compounds, (e.g., [Flavin group|flavin] and heme). Organic cofactors can be either prosthetic groups, which are tightly bound to an enzyme, or coenzymes, which are released from the enzyme's active site during the reaction.
Isoenzymes
[edit]Isoenzymes, or isozymes, are multiple forms of an enzyme, with slightly different protein sequence and closely similar but usually not identical functions. They are either products of different genes, or else different products of alternative splicing. They may either be produced in different organs or cell types to perform the same function, or several isoenzymes may be produced in the same cell type under differential regulation to suit the needs of changing development or environment. LDH (lactate dehydrogenase) has multiple isozymes, while fetal hemoglobin is an example of a developmentally regulated isoform of a non-enzymatic protein. The relative levels of isoenzymes in blood can be used to diagnose problems in the organ of secretion .
See also
[edit]References
[edit]- ^ Bunge, M. (1979). Treatise on Basic Philosophy, vol. 4. Ontology II: A World of Systems, p. 61-2. link.
- ^ Voon, C. H.; Sam, S. T. (2019). "2.1 Biosensors". Nanobiosensors for Biomolecular Targeting. Elsevier. ISBN 978-0-12-813900-4.
- ^ endogeny. (2011) Segen's Medical Dictionary. The Free Dictionary by Farlex. Farlex, Inc. Accessed June 27, 2019.
- ^ Green, D. E.; Goldberger, R. (1967). Molecular Insights into the Living Process. New York: Academic Press – via Google Books.
- ^ Gayon, J. (1998). "La philosophie et la biologie". In Mattéi, J. F. (ed.). Encyclopédie philosophique universelle. Vol. IV, Le Discours philosophique. Presses Universitaires de France. pp. 2152–2171. ISBN 9782130448631 – via Google Books.
- ^ a b c d Slabaugh, Michael R. & Seager, Spencer L. (2007). Organic and Biochemistry for Today (6th ed.). Pacific Grove: Brooks Cole. ISBN 978-0-495-11280-8.
- ^ a b Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Wlater P (2002). Molecular biology of the cell (4th ed.). New York: Garland Science. pp. 120–1. ISBN 0-8153-3218-1.
- ^ Saenger W (1984). Principles of Nucleic Acid Structure. Springer-Verlag. ISBN 0387907629.
- ^ Richardson JS, Schneider B, Murray LW, Kapral GJ, Immormino RM, Headd JJ, Richardson DC, Ham D, Hershkovits E, Williams LD, Keating KS, Pyle AM, Micallef D, Westbrook J, Berman HM (2008). "RNA Backbone: Consensus all-angle conformers and modular string nomenclature". RNA. 14 (3): 465–481. doi:10.1261/rna.657708. PMC 2248255. PMID 18192612.
- ^ Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR (1982). "Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena". Cell. 31 (1): 147–157. doi:10.1016/0092-8674(82)90414-7. PMID 6297745. S2CID 14787080.
- ^ a b Moran, Timothy H. (June 2009). "Fructose and Satiety". The Journal of Nutrition. 139 (6): 1253S–1256S. doi:10.3945/jn.108.097956. PMC 6459054. PMID 19403706.
- ^ Pigman, W.; D. Horton (1972). The Carbohydrates. Vol. 1A. San Diego: Academic Press. p. 3. ISBN 978-0-12-395934-8.
- ^ Jin, Tan; Wang He-Fang & Yan Xiu-Ping (2009). "Discrimination of Saccharides with a Fluorescent Molecular Imprinting Sensor Array Based on Phenylboronic Acid Functionalized Mesoporous Silica". Anal. Chem. 81 (13): 5273–80. doi:10.1021/ac900484x. PMID 19507843.
- ^ Bo Peng & Yu Qin (2008). "Lipophilic Polymer Membrane Optical Sensor with a Synthetic Receptor for Saccharide Detection". Anal. Chem. 80 (15): 6137–41. doi:10.1021/ac800946p. PMID 18593197.
- ^ K. Freudenberg; A.C. Nash, eds. (1968). Constitution and Biosynthesis of Lignin. Berlin: Springer-Verlag.
- ^ Richardson, JS (1981). "The Anatomy and Taxonomy of Proteins". Advances in Protein Chemistry. 34: 167–339. doi:10.1016/S0065-3233(08)60520-3. PMID 7020376. Archived from the original on 2019-03-16. Retrieved 2012-06-24.
External links
[edit]- Society for Biomolecular Sciences provider of a forum for education and information exchange among professionals within drug discovery and related disciplines.
