Microfilament
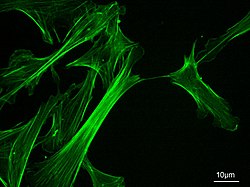
Microfilaments, also called actin filaments, are protein filaments in the cytoplasm of eukaryotic cells that form part of the cytoskeleton. They are primarily composed of polymers of actin, but are modified by and interact with numerous other proteins in the cell. Microfilaments are usually about 7 nm in diameter and made up of two strands of actin. Microfilament functions include cytokinesis, amoeboid movement, cell motility, changes in cell shape, endocytosis and exocytosis, cell contractility, and mechanical stability. Microfilaments are flexible and relatively strong, resisting buckling by multi-piconewton compressive forces and filament fracture by nanonewton tensile forces. In inducing cell motility, one end of the actin filament elongates while the other end contracts, presumably by myosin II molecular motors.[1] Additionally, they function as part of actomyosin-driven contractile molecular motors, wherein the thin filaments serve as tensile platforms for myosin's ATP-dependent pulling action in muscle contraction and pseudopod advancement. Microfilaments have a tough, flexible framework which helps the cell in movement.[2]
Actin was first discovered in rabbit skeletal muscle in the mid 1940s by F.B. Straub.[3] Almost 20 years later, H.E. Huxley demonstrated that actin is essential for muscle constriction. The mechanism in which actin creates long filaments was first described in the mid 1980s. Later studies showed that actin has an important role in cell shape, motility, and cytokinesis.
Organization
[edit]Actin filaments are assembled in two general types of structures: bundles and networks. Bundles can be composed of polar filament arrays, in which all barbed ends point to the same end of the bundle, or non-polar arrays, where the barbed ends point towards both ends. A class of actin-binding proteins, called cross-linking proteins, dictate the formation of these structures. Cross-linking proteins determine filament orientation and spacing in the bundles and networks. These structures are regulated by many other classes of actin-binding proteins, including motor proteins, branching proteins, severing proteins, polymerization promoters, and capping proteins.[citation needed]
In vitro self-assembly
[edit]Measuring approximately 6 nm in diameter,[4] microfilaments are the thinnest fibers of the cytoskeleton. They are polymers of actin subunits (globular actin, or G-actin), which as part of the fiber are referred to as filamentous actin, or F-actin. Each microfilament is made up of two helical, interlaced strands of subunits. Much like microtubules, actin filaments are polarized. Electron micrographs have provided evidence of their fast-growing barbed-ends and their slow-growing pointed-end. This polarity has been determined by the pattern created by the binding of myosin S1 fragments: they themselves are subunits of the larger myosin II protein complex. The pointed end is commonly referred to as the minus (−) end and the barbed end is referred to as the plus (+) end.[citation needed]
In vitro actin polymerization, or nucleation, starts with the self-association of three G-actin monomers to form a trimer. ATP-bound actin then itself binds the barbed end, and the ATP is subsequently hydrolyzed. ATP hydrolysis occurs with a half time of about 2 seconds,[5] while the half time for the dissociation of the inorganic phosphate is about 6 minutes.[5] This autocatalyzed event reduces the binding strength between neighboring subunits, and thus generally destabilizes the filament. In vivo actin polymerization is catalyzed by a class of filament end-tracking molecular motors known as actoclampins. Recent evidence suggests that the rate of ATP hydrolysis and the rate of monomer incorporation are strongly coupled.[citation needed]
Subsequently, ADP-actin dissociates slowly from the pointed end, a process significantly accelerated by the actin-binding protein, cofilin. ADP bound cofilin severs ADP-rich regions nearest the (−)-ends. Upon release, the free actin monomer slowly dissociates from ADP, which in turn rapidly binds to the free ATP diffusing in the cytosol, thereby forming the ATP-actin monomeric units needed for further barbed-end filament elongation. This rapid turnover is important for the cell's movement. End-capping proteins such as CapZ prevent the addition or loss of monomers at the filament end where actin turnover is unfavorable, such as in the muscle apparatus.[citation needed]
Actin polymerization together with capping proteins were recently used to control the 3-dimensional growth of protein filament so as to perform 3D topologies useful in technology and the making of electrical interconnect. Electrical conductivity is obtained by metallisation of the protein 3D structure.[6][7]
Mechanism of force generation
[edit]As a result of ATP hydrolysis, filaments elongate approximately 10 times faster at their barbed ends than their pointed ends. At steady-state, the polymerization rate at the barbed end matches the depolymerization rate at the pointed end, and microfilaments are said to be treadmilling. Treadmilling results in elongation in the barbed end and shortening in the pointed-end, so that the filament in total moves. Since both processes are energetically favorable, this means force is generated, the energy ultimately coming from ATP.[1]
Actin in cells
[edit]Intracellular actin cytoskeletal assembly and disassembly are tightly regulated by cell signaling mechanisms. Many signal transduction systems use the actin cytoskeleton as a scaffold, holding them at or near the inner face of the peripheral membrane. This subcellular location allows immediate responsiveness to transmembrane receptor action and the resulting cascade of signal-processing enzymes.[citation needed]
Because actin monomers must be recycled to sustain high rates of actin-based motility during chemotaxis, cell signalling is believed to activate cofilin, the actin-filament depolymerizing protein which binds to ADP-rich actin subunits nearest the filament's pointed-end and promotes filament fragmentation, with concomitant depolymerization in order to liberate actin monomers. In most animal cells, monomeric actin is bound to profilin and thymosin beta-4, both of which preferentially bind with one-to-one stoichiometry to ATP-containing monomers. Although thymosin beta-4 is strictly a monomer-sequestering protein, the behavior of profilin is far more complex. Profilin enhances the ability of monomers to assemble by stimulating the exchange of actin-bound ADP for solution-phase ATP to yield actin-ATP and ADP. Profilin is transferred to the leading edge by virtue of its PIP2 binding site, and it employs its poly-L-proline binding site to dock onto end-tracking proteins. Once bound, profilin-actin-ATP is loaded into the monomer-insertion site of actoclampin motors.[citation needed]
Another important component in filament formation is the Arp2/3 complex, which binds to the side of an already existing filament (or "mother filament"), where it nucleates the formation of a new daughter filament at a 70-degree angle relative to the mother filament, effecting a fan-like branched filament network.[8]
Specialized unique actin cytoskeletal structures are found adjacent to the plasma membrane. Four remarkable examples include red blood cells, human embryonic kidney cells, neurons, and sperm cells. In red blood cells, a spectrin-actin hexagonal lattice is formed by interconnected short actin filaments.[9] In human embryonic kidney cells, the cortical actin forms a scale-free fractal structure.[10] First found in neuronal axons, actin forms periodic rings that are stabilized by spectrin and adducin[11][12] – and this ring structure was then found by He et al 2016 to occur in almost every neuronal type and glial cells, across seemingly every animal taxon including Caenorhabditis elegans, Drosophila, Gallus gallus and Mus musculus.[13] And in mammalian sperm, actin forms a helical structure in the midpiece, i.e., the first segment of the flagellum.[14]
Associated proteins
[edit]In non-muscle cells, actin filaments are formed proximal to membrane surfaces. Their formation and turnover are regulated by many proteins, including:[citation needed]
- Filament end-tracking protein (e.g., formins, VASP, N-WASP)
- Filament-nucleator known as the Actin-Related Protein-2/3 (or Arp2/3) complex
- Filament cross-linkers (e.g., α-actinin, fascin, and fimbrin)
- Actin monomer-binding proteins profilin and thymosin β4
- Filament barbed-end cappers such as Capping Protein and CapG, etc.
- Filament-severing proteins like gelsolin.
- Actin depolymerizing proteins such as ADF/cofilin.
The actin filament network in non-muscle cells is highly dynamic. The actin filament network is arranged with the barbed-end of each filament attached to the cell's peripheral membrane by means of clamped-filament elongation motors, the above-mentioned "actoclampins", formed from a filament barbed-end and a clamping protein (formins, VASP, Mena, WASP, and N-WASP).[15] The primary substrate for these elongation motors is profilin-actin-ATP complex which is directly transferred to elongating filament ends.[16] The pointed-end of each filament is oriented toward the cell's interior. In the case of lamellipodial growth, the Arp2/3 complex generates a branched network, and in filopodia a parallel array of filaments is formed.[citation needed]
Actin acts as a track for myosin motor motility
[edit]Myosin motors are intracellular ATP-dependent enzymes that bind to and move along actin filaments. Various classes of myosin motors have very different behaviors, including exerting tension in the cell and transporting cargo vesicles.[citation needed]
A proposed model – actoclampins track filament ends
[edit]One proposed model suggests the existence of actin filament barbed-end-tracking molecular motors termed "actoclampin".[17] The proposed actoclampins generate the propulsive forces needed for actin-based motility of lamellipodia, filopodia, invadipodia, dendritic spines, intracellular vesicles, and motile processes in endocytosis, exocytosis, podosome formation, and phagocytosis. Actoclampin motors also propel such intracellular pathogens as Listeria monocytogenes, Shigella flexneri, Vaccinia and Rickettsia. When assembled under suitable conditions, these end-tracking molecular motors can also propel biomimetic particles.[citation needed]
The term actoclampin is derived from acto- to indicate the involvement of an actin filament, as in actomyosin, and clamp to indicate a clasping device used for strengthening flexible/moving objects and for securely fastening two or more components, followed by the suffix -in to indicate its protein origin. An actin filament end-tracking protein may thus be termed a clampin.[citation needed]
Dickinson and Purich recognized that prompt ATP hydrolysis could explain the forces achieved during actin-based motility.[15] They proposed a simple mechanoenzymatic sequence known as the Lock, Load & Fire Model, in which an end-tracking protein remains tightly bound ("locked" or clamped) onto the end of one sub-filament of the double-stranded actin filament. After binding to Glycyl-Prolyl-Prolyl-Prolyl-Prolyl-Prolyl-registers on tracker proteins, Profilin-ATP-actin is delivered ("loaded") to the unclamped end of the other sub-filament, whereupon ATP within the already clamped terminal subunit of the other subfragment is hydrolyzed ("fired"), providing the energy needed to release that arm of the end-tracker, which then can bind another Profilin-ATP-actin to begin a new monomer-addition round.[citation needed]
Steps involved
[edit]The following steps describe one force-generating cycle of an actoclampin molecular motor:
- The polymerization cofactor profilin and the ATP·actin combine to form a profilin-ATP-actin complex that then binds to the end-tracking unit
- The cofactor and monomer are transferred to the barbed-end of an actin already clamped filament
- The tracking unit and cofactor dissociate from the adjacent protofilament, in a step that can be facilitated by ATP hydrolysis energy to modulate the affinity of the cofactor and/or the tracking unit for the filament; and this mechanoenzymatic cycle is then repeated, starting this time on the other sub-filament growth site.[citation needed]
When operating with the benefit of ATP hydrolysis, AC motors generate per-filament forces of 8–9 pN, which is far greater than the per-filament limit of 1–2 pN for motors operating without ATP hydrolysis.[15][17][18] The term actoclampin is generic and applies to all actin filament end-tracking molecular motors, irrespective of whether they are driven actively by an ATP-activated mechanism or passively.[citation needed]
Some actoclampins (e.g., those involving Ena/VASP proteins, WASP, and N-WASP) apparently require Arp2/3-mediated filament initiation to form the actin polymerization nucleus that is then "loaded" onto the end-tracker before processive motility can commence. To generate a new filament, Arp2/3 requires a "mother" filament, monomeric ATP-actin, and an activating domain from Listeria ActA or the VCA region of N-WASP. The Arp2/3 complex binds to the side of the mother filament, forming a Y-shaped branch having a 70-degree angle with respect to the longitudinal axis of the mother filament. Then upon activation by ActA or VCA, the Arp complex is believed to undergo a major conformational change, bringing its two actin-related protein subunits near enough to each other to generate a new filament gate. Whether ATP hydrolysis may be required for nucleation and/or Y-branch release is a matter under active investigation.[citation needed]
References
[edit]- ^ a b Roberts K, Raff M, Alberts B, Walter P, Lewis J, Johnson A (March 2002). Molecular Biology of the Cell (4th ed.). Routledge. p. 1616. ISBN 0-8153-3218-1.
- ^ Gunning PW, Ghoshdastider U, Whitaker S, Popp D, Robinson RC (June 2015). "The evolution of compositionally and functionally distinct actin filaments". Journal of Cell Science. 128 (11): 2009–19. doi:10.1242/jcs.165563. PMID 25788699.
- ^ Kelpsch, Daniel J.; Tootle, Tina L. (December 2018). "Nuclear actin: from discovery to function". Anatomical Record. 301 (12): 1999–2013. doi:10.1002/ar.23959. ISSN 1932-8486. PMC 6289869. PMID 30312531.
- ^ Fuchs E, Cleveland DW (January 1998). "A structural scaffolding of intermediate filaments in health and disease". Science. 279 (5350): 514–9. Bibcode:1998Sci...279..514F. doi:10.1126/science.279.5350.514. PMID 9438837.
- ^ a b Pollard TD, Earnshaw WD (2004). Cell Biology (First ed.). SAUNDERS. ISBN 978-1-4160-2388-3.
- ^ Galland R, Leduc P, Guérin C, Peyrade D, Blanchoin L, Théry M (May 2013). "Fabrication of three-dimensional electrical connections by means of directed actin self-organization". Nature Materials. 12 (5): 416–21. Bibcode:2013NatMa..12..416G. doi:10.1038/nmat3569. PMID 23396247.
- ^ US Patent US 9070702, Method for obtaining three-dimensional actin structures and uses thereof, Jean-Christophe Gabriel, Laurent Blanchoin, Manuel Thery, Remi Galland
- ^ Mullins RD, Heuser JA, Pollard TD (May 1998). "The interaction of Arp2/3 complex with actin: nucleation, high affinity pointed end capping, and formation of branching networks of filaments". Proceedings of the National Academy of Sciences of the United States of America. 95 (11): 6181–6. Bibcode:1998PNAS...95.6181M. doi:10.1073/pnas.95.11.6181. PMC 27619. PMID 9600938.
- ^ Gokhin, David S.; Fowler, Velia M. (May 2016). "Feisty filaments: actin dynamics in the red blood cell membrane skeleton". Current Opinion in Hematology. 23 (3): 206–214. doi:10.1097/MOH.0000000000000227. ISSN 1065-6251. PMC 4966542. PMID 27055045.
- ^ Sadegh, Sanaz; Higgins, Jenny L.; Mannion, Patrick C.; Tamkun, Michael M.; Krapf, Diego (2017-03-09). "Plasma Membrane is Compartmentalized by a Self-Similar Cortical Actin Meshwork". Physical Review X. 7 (1): 011031. arXiv:1702.03997. Bibcode:2017PhRvX...7a1031S. doi:10.1103/PhysRevX.7.011031. ISSN 2160-3308. PMC 5500227. PMID 28690919.
- ^ Xu, K.; Zhong, G.; Zhuang, X. (2013-01-25). "Actin, Spectrin, and Associated Proteins Form a Periodic Cytoskeletal Structure in Axons". Science. 339 (6118): 452–456. Bibcode:2013Sci...339..452X. doi:10.1126/science.1232251. ISSN 0036-8075. PMC 3815867. PMID 23239625.
- ^ D'Este, Elisa; Kamin, Dirk; Göttfert, Fabian; El-Hady, Ahmed; Hell, Stefan W. (March 2015). "STED Nanoscopy Reveals the Ubiquity of Subcortical Cytoskeleton Periodicity in Living Neurons". Cell Reports. 10 (8): 1246–1251. doi:10.1016/j.celrep.2015.02.007. hdl:11858/00-001M-0000-0025-05F3-D. PMID 25732815.
- ^ Sahl, Steffen J.; Hell, Stefan W.; Jakobs, Stefan (2017-09-06). "Fluorescence nanoscopy in cell biology". Nature Reviews Molecular Cell Biology. 18 (11). Nature Portfolio: 685–701. doi:10.1038/nrm.2017.71. ISSN 1471-0072. PMID 28875992. S2CID 20747438.
- ^ Gervasi, María G.; Xu, Xinran; Carbajal-Gonzalez, Blanca; Buffone, Mariano G.; Visconti, Pablo E.; Krapf, Diego (2018-06-01). "The actin cytoskeleton of the mouse sperm flagellum is organized in a helical structure". Journal of Cell Science. 131 (11): jcs215897. doi:10.1242/jcs.215897. ISSN 0021-9533. PMC 6031324. PMID 29739876.
- ^ a b c Dickinson RB, Purich DL (January 2007). "Nematode sperm motility: nonpolar filament polymerization mediated by end-tracking motors". Biophysical Journal. 92 (2): 622–31. Bibcode:2007BpJ....92..622D. doi:10.1529/biophysj.106.090472. PMC 1751402. PMID 17056726.
- ^ Dickinson RB, Southwick FS, Purich DL (October 2002). "A direct-transfer polymerization model explains how the multiple profilin-binding sites in the actoclampin motor promote rapid actin-based motility". Archives of Biochemistry and Biophysics. 406 (2): 296–301. doi:10.1016/s0003-9861(02)00212-6. PMID 12361718.
- ^ a b Dickinson RB, Caro L, Purich DL (October 2004). "Force generation by cytoskeletal filament end-tracking proteins". Biophysical Journal. 87 (4): 2838–54. Bibcode:2004BpJ....87.2838D. doi:10.1529/biophysj.104.045211. PMC 1304702. PMID 15454475.
- ^ Dickinson RB, Purich DL (August 2006). "Diffusion rate limitations in actin-based propulsion of hard and deformable particles". Biophysical Journal. 91 (4): 1548–63. Bibcode:2006BpJ....91.1548D. doi:10.1529/biophysj.106.082362. PMC 1518650. PMID 16731556.
External links
[edit]- Microfilaments at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Microfilament+proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- "Microfilament" at Dorland's Medical Dictionary
