Senecavirus
| Senecavirus | |
|---|---|
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Picornavirales |
| Family: | Picornaviridae |
| Genus: | Senecavirus |
| Synonyms | |
| |
Senecavirus is a genus of viruses in the order Picornavirales, in the family Picornaviridae. Mammals of the order Artiodactyla (even-toed ungulates) serve as natural hosts, with senecaviruses reported in cows, pigs and dolphins. There is only one species in this genus: Senecavirus A.[1][2] Senecavirus is a replication-competent oncolytic picornavirus. It has selective tropism for cancers with neuroendocrine features including small cell lung cancer (SCLC) and several pediatric solid tumors including retinoblastoma, neuroblastoma, and medulloblastoma.[3] A Phase I clinical trial of Senecavirus in adults with neuroendocrine tumors showed that senecavirus is apparently safe to administer at doses up to 1E11 vp/kg.[4] It has potential antineoplastic activity.[5][6]
Structure
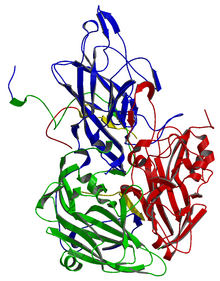
[edit]Viruses in Senecavirus are non-enveloped, with icosahedral, spherical, and round geometries, with T=pseudo3 symmetry. The diameter is around 30 nm. Genomes are linear and non-segmented, around 7.3kb in length.[1]
| Genus | Structure | Symmetry | Capsid | Genomic arrangement | Genomic segmentation |
|---|---|---|---|---|---|
| Senecavirus | Icosahedral | Pseudo T=3 | Non-enveloped | Linear | Monopartite |
Life cycle
[edit]Viral replication is cytoplasmic. Entry into the host cell is achieved by attachment of the virus to host receptors, which mediates endocytosis. Replication follows the positive stranded RNA virus replication model. Positive stranded RNA virus transcription is the method of transcription. The virus exits the host cell by lysis, and viroporins. Pig and maybe also cow serve as the natural host.[1]
The receptor for Seneca Valley virus has been identified as anthrax toxin receptor 1.[7]
| Genus | Host details | Tissue tropism | Entry details | Release details | Replication site | Assembly site | Transmission |
|---|---|---|---|---|---|---|---|
| Senecavirus | Pigs, Cow | Oncolytic | Cell receptor endocytosis | Lysis | Cytoplasm | Cytoplasm | Unknown |
Discovery and origin
[edit]The complete genome sequence of senecavirus was completed in 2008.[8]
An infectious clone of senecavirus was reported in 2012.[9]
Senecavirus has been proposed to attack cancer stem cells.[10]
Diagnostic monoclonal antibodies have been generated against senecavirus.[11]
While the sequence of SVV's protein-coding genome is most similar to members in the Cardiovirus genus, the non-coding RNA internal ribosome entry site (IRES) is most similar to those of the Pestivirus genus, including classical swine fever virus, and Hepacivirus genus, including Hepatitis C virus.[12]
The SVV IRES RNA shares similarities in sequence, structure, and function with the hepatitis C virus IRES. Subdomain IIa of the SVV and HCV IRES shares a similar structure and ligand-binding function as seen in its crystal structure.[13] This subdomain IIa region is classified as a ligand-responsive RNA switch which adopts well-defined ligand-free and bound conformations without breaking or forming any base pairs in its secondary structure upon interconversion between the two states.[14] This RNA switch from the SVV IRES has been incorporated into triangular RNA nanostructures.[15]
Clinical trials
[edit]The initial isolate is being developed as an anti-cancer therapeutic by virtual company Neotropix, Inc. under the name NTX-010.
Phase I
- Safety study of senecavirus in patients with solid tumors with neuroendocrine features.[16] This study was published in 2011 and the data show that the virus was well tolerated by 30 patients and some signs of anti-tumour activity were observed. The data warranted further investigation of the virus in a phase II trial in small cell lung cancer.[4]
Phase II
- Senecavirus after chemotherapy in treating patients with extensive-stage small cell lung cancer[17]
- Senecavirus and cyclophosphamide in young patients with neuroblastoma, rhabdomyosarcoma, or rare tumors with neuroendocrine features[18]
Virus replication
[edit]Senecavirus uses the anthrax toxin receptor 1 (ANTXR1) protein as a receptor.[19] A high-resolution structure of senecavirus with this receptor has been published.[20]
See also
[edit]References
[edit]- ^ a b c "Viral Zone". ExPASy. Retrieved 15 June 2015.
- ^ "Virus Taxonomy: 2020 Release". International Committee on Taxonomy of Viruses (ICTV). March 2021. Retrieved 20 May 2021.
- ^ Reddy PS, Burroughs KD, Hales LM, Ganesh S, Jones BH, Idamakanti N, Hay C, Li SS, Skele KL, Vasko AJ, Yang J, Watkins DN, Rudin CM, Hallenbeck PL (2007). "Seneca Valley Virus, a Systemically Deliverable Oncolytic Picornavirus, and the Treatment of Neuroendocrine Cancers". JNCI Journal of the National Cancer Institute. 99 (21): 1623–1633. doi:10.1093/jnci/djm198. PMC 5261858. PMID 17971529.
- ^ a b Rudin CM, Poirier JT, Senzer NN, Stephenson J, Loesch D, Burroughs KD, Reddy PS, Hann CL, Hallenbeck PL (February 15, 2011). "Phase I clinical study of Seneca Valley Virus (SVV-001), a replication-competent picornavirus, in advanced solid tumors with neuroendocrine features". Clinical Cancer Research. 17 (4): 888–95. doi:10.1158/1078-0432.CCR-10-1706. PMC 5317273. PMID 21304001.
- ^ National Cancer Institute Definition of Seneca Valley virus-001. National Cancer Institute Retrieved on 2008-10-09.
- ^ Morton CL, Houghton PJ, Kolb EA, et al. (August 2010). "Initial testing of the replication competent Seneca Valley virus (NTX-010) by the pediatric preclinical testing program". Pediatr Blood Cancer. 55 (2): 295–303. doi:10.1002/pbc.22535. PMC 3003870. PMID 20582972.
- ^ Miles LA, Burga LN, Gardner EE, Bostina M, Poirier JT, Rudin CM (2017) Anthrax toxin receptor 1 is the cellular receptor for Seneca Valley virus. J Clin Invest
- ^ Hales LM, Knowles NJ, Reddy PS, Xu L, Hay C, Hallenbeck PL (2008). "Complete genome sequence analysis of Seneca Valley virus-001, a novel oncolytic picornavirus". Journal of General Virology. 89 (5): 1265–1275. doi:10.1099/vir.0.83570-0. PMID 18420805.
- ^ Poirier JT, Reddy PS, Idamakanti N, Li SS, Stump KL, Burroughs KD, Hallenbeck PL, Rudin CM (2012). "Characterization of a full-length infectious cDNA clone and a GFP reporter derivative of the oncolytic picornavirus SVV-001". Journal of General Virology. 93 (Pt 12): 2606–2613. doi:10.1099/vir.0.046011-0. PMID 22971818.
- ^ Friedman GK, Cassady KA, Beierle EA, Markert JM, Gillespie GY (2012). "Targeting pediatric cancer stem cells with oncolytic virotherapy". Pediatric Research. 71 (4–2): 500–510. doi:10.1038/pr.2011.58. PMC 3607376. PMID 22430386.
- ^ Yang M, Van Bruggen R, Xu W (2011). "Generation and diagnostic application of monoclonal antibodies against Seneca Valley virus". Journal of Veterinary Diagnostic Investigation. 24 (1): 42–50. doi:10.1177/1040638711426323. PMID 22362934.
- ^ Willcocks MM, Locker N, Gomwalk Z, Royall E, Bakhshesh M, Belsham GJ, Idamakanti N, Burroughs KD, Reddy PS, Hallenbeck PL, Roberts LO (2011). "Structural Features of the Seneca Valley Virus Internal Ribosome Entry Site (IRES) Element: A Picornavirus with a Pestivirus-Like IRES". Journal of Virology. 85 (9): 4452–4461. doi:10.1128/JVI.01107-10. PMC 3126232. PMID 21325406.
- ^ Boerneke M, Dibrov S, Gu J, Wyles D, Hermann T (November 11, 2014). "Functional conservation despite structural divergence in ligand-responsive RNA switches". Proc Natl Acad Sci U S A. 111 (45): 15952–7. Bibcode:2014PNAS..11115952B. doi:10.1073/pnas.1414678111. PMC 4234586. PMID 25349403.
- ^ Boerneke M, Hermann T (August 2015). "Ligand-responsive RNA mechanical switches". RNA Biology. 12 (8): 780–786. doi:10.1080/15476286.2015.1054592. PMC 4615790. PMID 26158858.
- ^ Boerneke M, Dibrov S, Hermann T (February 23, 2016). "Crystal-Structure-Guided Design of Self-Assembling RNA Nanotriangles". Angew Chem Int Ed Engl. 55 (12): 4097–100. doi:10.1002/anie.201600233. PMC 4824544. PMID 26914842.
- ^ "Safety Study of Seneca Valley Virus in Patients With Solid Tumors With Neuroendocrine Features". ClinicalTrials.gov. 23 February 2010.
- ^ "Seneca Valley Virus-001 After Chemotherapy in Treating Patients With Extensive-Stage Small Cell Lung Cancer". clinicaltrials.gov.
- ^ "Seneca Valley Virus-001 and Cyclophosphamide in Treating Young Patients With Relapsed or Refractory Neuroblastoma, Rhabdomyosarcoma, or Rare Tumors With Neuroendocrine Features". clinicaltrials.gov.
- ^ Miles LA, Burga LN, Gardner EE, Bostina M, Poirier JT, Rudin CM (1 August 2017). "Anthrax toxin receptor 1 is the cellular receptor for Seneca Valley virus". The Journal of Clinical Investigation. 127 (8): 2957–2967. doi:10.1172/JCI93472. PMC 5531414. PMID 28650343.
- ^ Jayawardena N, Burga LN, Easingwood RA, Takizawa Y, Wolf M, Bostina M (13 November 2018). "Structural basis for anthrax toxin receptor 1 recognition by Seneca Valley Virus". Proceedings of the National Academy of Sciences of the United States of America. 115 (46): E10934–E10940. Bibcode:2018PNAS..11510934J. doi:10.1073/pnas.1810664115. PMC 6243253. PMID 30381454.
External links
[edit]- Structure of Seneca Valley Virus-001 — on Virus Particle ExploreR (VIPERdb)
- Viralzone: Senecavirus
- ICTV
