Non-covalent interaction
It has been suggested that this article be merged with Supramolecular chemistry. (Discuss) Proposed since October 2024. |
In chemistry, a non-covalent interaction differs from a covalent bond in that it does not involve the sharing of electrons,[1] but rather involves more dispersed variations of electromagnetic interactions between molecules or within a molecule. The chemical energy released in the formation of non-covalent interactions is typically on the order of 1–5 kcal/mol (1000–5000 calories per 6.02×1023 molecules).[2] Non-covalent interactions can be classified into different categories, such as electrostatic, π-effects, van der Waals forces, and hydrophobic effects.[3][2]
Non-covalent interactions[4] are critical in maintaining the three-dimensional structure of large molecules, such as proteins and nucleic acids. They are also involved in many biological processes in which large molecules bind specifically but transiently to one another (see the properties section of the DNA page). These interactions also heavily influence drug design, crystallinity and design of materials, particularly for self-assembly, and, in general, the synthesis of many organic molecules.[3][5][6][7][8]
The non-covalent interactions may occur between different parts of the same molecule (e.g. during protein folding) or between different molecules and therefore are discussed also as intermolecular forces.
Electrostatic interactions
[edit]Ionic
[edit]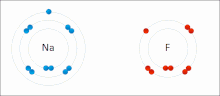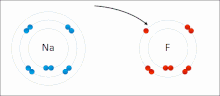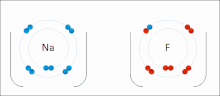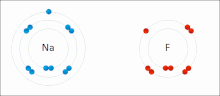
Ionic interactions involve the attraction of ions or molecules with full permanent charges of opposite signs. For example, sodium fluoride involves the attraction of the positive charge on sodium (Na+) with the negative charge on fluoride (F−).[9] However, this particular interaction is easily broken upon addition to water, or other highly polar solvents. In water ion pairing is mostly entropy driven; a single salt bridge usually amounts to an attraction value of about ΔG =5 kJ/mol at an intermediate ion strength I, at I close to zero the value increases to about 8 kJ/mol. The ΔG values are usually additive and largely independent of the nature of the participating ions, except for transition metal ions etc.[10]
These interactions can also be seen in molecules with a localized charge on a particular atom. For example, the full negative charge associated with ethoxide, the conjugate base of ethanol, is most commonly accompanied by the positive charge of an alkali metal salt such as the sodium cation (Na+).
Hydrogen bonding
[edit]
A hydrogen bond (H-bond), is a specific type of interaction that involves dipole–dipole attraction between a partially positive hydrogen atom and a highly electronegative, partially negative oxygen, nitrogen, sulfur, or fluorine atom (not covalently bound to said hydrogen atom). It is not a covalent bond, but instead is classified as a strong non-covalent interaction. It is responsible for why water is a liquid at room temperature and not a gas (given water's low molecular weight). Most commonly, the strength of hydrogen bonds lies between 0–4 kcal/mol, but can sometimes be as strong as 40 kcal/mol[3] In solvents such as chloroform or carbon tetrachloride one observes e.g. for the interaction between amides additive values of about 5 kJ/mol. According to Linus Pauling the strength of a hydrogen bond is essentially determined by the electrostatic charges. Measurements of thousands of complexes in chloroform or carbon tetrachloride have led to additive free energy increments for all kind of donor-acceptor combinations.[11][12]
Halogen bonding
[edit]
Halogen bonding is a type of non-covalent interaction which does not involve the formation nor breaking of actual bonds, but rather is similar to the dipole–dipole interaction known as hydrogen bonding. In halogen bonding, a halogen atom acts as an electrophile, or electron-seeking species, and forms a weak electrostatic interaction with a nucleophile, or electron-rich species. The nucleophilic agent in these interactions tends to be highly electronegative (such as oxygen, nitrogen, or sulfur), or may be anionic, bearing a negative formal charge. As compared to hydrogen bonding, the halogen atom takes the place of the partially positively charged hydrogen as the electrophile.[citation needed]
Halogen bonding should not be confused with halogen–aromatic interactions, as the two are related but differ by definition. Halogen–aromatic interactions involve an electron-rich aromatic π-cloud as a nucleophile; halogen bonding is restricted to monatomic nucleophiles.[5]
Van der Waals forces
[edit]Van der Waals forces are a subset of electrostatic interactions involving permanent or induced dipoles (or multipoles). These include the following:
- permanent dipole–dipole interactions, alternatively called the Keesom force
- dipole-induced dipole interactions, or the Debye force
- induced dipole-induced dipole interactions, commonly referred to as London dispersion forces
Hydrogen bonding and halogen bonding are typically not classified as Van der Waals forces.
Dipole–dipole
[edit]
Dipole-dipole interactions are electrostatic interactions between permanent dipoles in molecules. These interactions tend to align the molecules to increase attraction (reducing potential energy). Normally, dipoles are associated with electronegative atoms, including oxygen, nitrogen, sulfur, and fluorine.
For example, acetone, the active ingredient in some nail polish removers, has a net dipole associated with the carbonyl (see figure 2). Since oxygen is more electronegative than the carbon that is covalently bonded to it, the electrons associated with that bond will be closer to the oxygen than the carbon, creating a partial negative charge (δ−) on the oxygen, and a partial positive charge (δ+) on the carbon. They are not full charges because the electrons are still shared through a covalent bond between the oxygen and carbon. If the electrons were no longer being shared, then the oxygen-carbon bond would be an electrostatic interaction.
Often molecules contain dipolar groups, but have no overall dipole moment. This occurs if there is symmetry within the molecule that causes the dipoles to cancel each other out. This occurs in molecules such as tetrachloromethane. Note that the dipole-dipole interaction between two individual atoms is usually zero, since atoms rarely carry a permanent dipole. See atomic dipoles.
Dipole-induced dipole
[edit]A dipole-induced dipole interaction (Debye force) is due to the approach of a molecule with a permanent dipole to another non-polar molecule with no permanent dipole. This approach causes the electrons of the non-polar molecule to be polarized toward or away from the dipole (or "induce" a dipole) of the approaching molecule.[13] Specifically, the dipole can cause electrostatic attraction or repulsion of the electrons from the non-polar molecule, depending on orientation of the incoming dipole.[13] Atoms with larger atomic radii are considered more "polarizable" and therefore experience greater attractions as a result of the Debye force.[citation needed]
London dispersion forces
[edit]London dispersion forces[14][15][16][17] are the weakest type of non-covalent interaction. In organic molecules, however, the multitude of contacts can lead to larger contributions, particularly in the presence of heteroatoms. They are also known as "induced dipole-induced dipole interactions" and present between all molecules, even those which inherently do not have permanent dipoles. Dispersive interactions increase with the polarizability of interacting groups, but are weakened by solvents of increased polarizability.[18] They are caused by the temporary repulsion of electrons away from the electrons of a neighboring molecule, leading to a partially positive dipole on one molecule and a partially negative dipole on another molecule.[6] Hexane is a good example of a molecule with no polarity or highly electronegative atoms, yet is a liquid at room temperature due mainly to London dispersion forces. In this example, when one hexane molecule approaches another, a temporary, weak partially negative dipole on the incoming hexane can polarize the electron cloud of another, causing a partially positive dipole on that hexane molecule. In absence of solvents hydrocarbons such as hexane form crystals due to dispersive forces ; the sublimation heat of crystals is a measure of the dispersive interaction. While these interactions are short-lived and very weak, they can be responsible for why certain non-polar molecules are liquids at room temperature.
π-effects
[edit]π-effects can be broken down into numerous categories, including π-stacking, cation-π and anion-π interactions, and polar-π interactions. In general, π-effects are associated with the interactions of molecules with the π-systems of arenes.[3]
π–π interaction
[edit]
π–π interactions are associated with the interaction between the π-orbitals of a molecular system.[3] The high polarizability of aromatic rings lead to dispersive interactions as major contribution to so-called stacking effects. These play a major role for interactions of nucleobases e.g. in DNA.[19] For a simple example, a benzene ring, with its fully conjugated π cloud, will interact in two major ways (and one minor way) with a neighboring benzene ring through a π–π interaction (see figure 3). The two major ways that benzene stacks are edge-to-face, with an enthalpy of ~2 kcal/mol, and displaced (or slip stacked), with an enthalpy of ~2.3 kcal/mol.[3] The sandwich configuration is not nearly as stable of an interaction as the previously two mentioned due to high electrostatic repulsion of the electrons in the π orbitals.[3]
Cation–π and anion–π interaction
[edit]
Cation–pi interactions can be as strong or stronger than H-bonding in some contexts.[3][20]
Anion–π interactions are very similar to cation–π interactions, but reversed. In this case, an anion sits atop an electron-poor π-system, usually established by the presence of electron-withdrawing substituents on the conjugated molecule[21]

Polar–π
[edit]Polar–π interactions involve molecules with permanent dipoles (such as water) interacting with the quadrupole moment of a π-system (such as that in benzene (see figure 5). While not as strong as a cation-π interaction, these interactions can be quite strong (~1-2 kcal/mol), and are commonly involved in protein folding and crystallinity of solids containing both hydrogen bonding and π-systems.[3] In fact, any molecule with a hydrogen bond donor (hydrogen bound to a highly electronegative atom) will have favorable electrostatic interactions with the electron-rich π-system of a conjugated molecule.[citation needed]
Hydrophobic effect
[edit]The hydrophobic effect is the desire for non-polar molecules to aggregate in aqueous solutions in order to separate from water.[22] This phenomenon leads to minimum exposed surface area of non-polar molecules to the polar water molecules (typically spherical droplets), and is commonly used in biochemistry to study protein folding and other various biological phenomenon.[22] The effect is also commonly seen when mixing various oils (including cooking oil) and water. Over time, oil sitting on top of water will begin to aggregate into large flattened spheres from smaller droplets, eventually leading to a film of all oil sitting atop a pool of water. However the hydrophobic effect is not considered a non-covalent interaction as it is a function of entropy and not a specific interaction between two molecules, usually characterized by entropy.enthalpy compensation.[23][24][25] An essentially enthalpic hydrophobic effect materializes if a limited number of water molecules are restricted within a cavity; displacement of such water molecules by a ligand frees the water molecules which then in the bulk water enjoy a maximum of hydrogen bonds close to four.[26][27]
Examples
[edit]Drug design
[edit]Most pharmaceutical drugs are small molecules which elicit a physiological response by "binding" to enzymes or receptors, causing an increase or decrease in the enzyme's ability to function. The binding of a small molecule to a protein is governed by a combination of steric, or spatial considerations, in addition to various non-covalent interactions, although some drugs do covalently modify an active site (see irreversible inhibitors). Using the "lock and key model" of enzyme binding, a drug (key) must be of roughly the proper dimensions to fit the enzyme's binding site (lock).[28] Using the appropriately sized molecular scaffold, drugs must also interact with the enzyme non-covalently in order to maximize binding affinity binding constant and reduce the ability of the drug to dissociate from the binding site. This is achieved by forming various non-covalent interactions between the small molecule and amino acids in the binding site, including: hydrogen bonding, electrostatic interactions, pi stacking, van der Waals interactions, and dipole–dipole interactions.
Non-covalent metallo drugs have been developed. For example, dinuclear triple-helical compounds in which three ligand strands wrap around two metals, resulting in a roughly cylindrical tetracation have been prepared. These compounds bind to the less-common nucleic acid structures, such as duplex DNA, Y-shaped fork structures and 4-way junctions.[29]
Protein folding and structure
[edit]The folding of proteins from a primary (linear) sequence of amino acids to a three-dimensional structure is directed by all types of non-covalent interactions, including the hydrophobic forces and formation of intramolecular hydrogen bonds. Three-dimensional structures of proteins, including the secondary and tertiary structures, are stabilized by formation of hydrogen bonds. Through a series of small conformational changes, spatial orientations are modified so as to arrive at the most energetically minimized orientation achievable. The folding of proteins is often facilitated by enzymes known as molecular chaperones.[30] Sterics, bond strain, and angle strain also play major roles in the folding of a protein from its primary sequence to its tertiary structure.
Single tertiary protein structures can also assemble to form protein complexes composed of multiple independently folded subunits. As a whole, this is called a protein's quaternary structure. The quaternary structure is generated by the formation of relatively strong non-covalent interactions, such as hydrogen bonds, between different subunits to generate a functional polymeric enzyme.[31] Some proteins also utilize non-covalent interactions to bind cofactors in the active site during catalysis, however a cofactor can also be covalently attached to an enzyme. Cofactors can be either organic or inorganic molecules which assist in the catalytic mechanism of the active enzyme. The strength with which a cofactor is bound to an enzyme may vary greatly; non-covalently bound cofactors are typically anchored by hydrogen bonds or electrostatic interactions.
Boiling points
[edit]Non-covalent interactions have a significant effect on the boiling point of a liquid. Boiling point is defined as the temperature at which the vapor pressure of a liquid is equal to the pressure surrounding the liquid. More simply, it is the temperature at which a liquid becomes a gas. As one might expect, the stronger the non-covalent interactions present for a substance, the higher its boiling point. For example, consider three compounds of similar chemical composition: sodium n-butoxide (C4H9ONa), diethyl ether (C4H10O), and n-butanol (C4H9OH).

The predominant non-covalent interactions associated with each species in solution are listed in the above figure. As previously discussed, ionic interactions require considerably more energy to break than hydrogen bonds, which in turn are require more energy than dipole–dipole interactions. The trends observed in their boiling points (figure 8) shows exactly the correlation expected, where sodium n-butoxide requires significantly more heat energy (higher temperature) to boil than n-butanol, which boils at a much higher temperature than diethyl ether. The heat energy required for a compound to change from liquid to gas is associated with the energy required to break the intermolecular forces each molecule experiences in its liquid state.
References
[edit]- ^ Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell J (2000). "Glossary". Molecular Cell Biology (4th ed.). New York: W.H. Freeman. ISBN 978-0-7167-3136-8.
- ^ a b Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell J (2000). "Noncovalent bonds". Molecular Cell Biology (4th ed.). New York: W.H. Freeman. ISBN 978-0-7167-3136-8.
- ^ a b c d e f g h i Anslyn E (2004). Modern Physical Organic Chemistry. Sausalito, CA: University Science. ISBN 978-1-891389-31-3.
- ^ Schalley CA (March 2012). "Introduction" (PDF). Analytical Methods in Supramolecular Chemistry (2nd ed.). Wiley. ISBN 978-3-527-32982-3.
- ^ a b Cockroft SL, Hunter CA (February 2007). "Chemical double-mutant cycles: dissecting non-covalent interactions". Chemical Society Reviews. 36 (2): 172–188. doi:10.1039/b603842p. PMID 17264921.
- ^ a b Brown TL, Bursten BE, Eugene H, LeMay H (2009). Chemistry: The Central Science (11th ed.). Upper Saddle River, NJ: Pearson Prentice Hall. ISBN 978-0-13-600617-6.
- ^ Eisler M (2010). "Self-Assembly". Encyclopedia of nanoscience and society. Thousand Oaks, Calif.: Sage. doi:10.4135/9781412972093.n412. ISBN 978-1-4129-7209-3.
- ^ Biedermann F, Schneider HJ (May 2016). "Experimental Binding Energies in Supramolecular Complexes". Chemical Reviews. 116 (9): 5216–5300. doi:10.1021/acs.chemrev.5b00583. PMID 27136957.
- ^ Ciferri A, Perico A, eds. (2012). Ionic Interactions in Natural and Synthetic Macromolecules. John Wiley & Sons. ISBN 978-0-470-52927-0.
- ^ Schneider HJ (2009). "Binding mechanisms in supramolecular complexes". Angewandte Chemie. 48 (22): 3924–3977. doi:10.1002/anie.200802947. PMID 19415701.
- ^ Abraham MH (1993). "Scales of Solute Hydrogen-bonding: their Construction and Application to Physicochemical and Biochemical Processes". Chem. Soc. Rev. 22 (2): 73–83. doi:10.1039/CS9932200073.
- ^ Raevsky OA, Skvortsov VS (June 2005). "Quantifying hydrogen bonding in QSAR and molecular modeling". SAR and QSAR in Environmental Research. 16 (3): 287–300. Bibcode:2005SQER...16..287R. doi:10.1080/10659360500036893. PMID 15804815. S2CID 34132761.
- ^ a b "Induced-Dipole Forces". Retrieved 11 November 2013.
- ^ Scheiner S, ed. (2015). Noncovalent Forces. Springer. ISBN 978-3-319-14162-6.
- ^ Otero de la Roza A, DiLabio GA, eds. (2017). Non-covalent Interactions in Quantum Chemistry and Physics: Theory and Applications. Elsevier. ISBN 978-0-12-809835-6.
- ^ Maharramov AM, Mahmudov KT, Kopylovich MN, Pombeiro AJ (2016). Non-covalent Interactions in the Synthesis and Design of New Compounds. Wiley. ISBN 978-1-119-10989-1.
- ^ Hobza P, Müller-Dethlefs K (2009). Non-Covalent Interactions: Theory and Experiment. Theoretical and Computational Chemistry Series. Royal Society of Chemistry. ISBN 978-1-84755-853-4.
- ^ Schneider HJ (July 2015). "Dispersive interactions in solution complexes". Accounts of Chemical Research. 48 (7): 1815–1822. doi:10.1021/acs.accounts.5b00111. PMID 26083908.
- ^ Riley KE, Hobza P (April 2013). "On the importance and origin of aromatic interactions in chemistry and biodisciplines". Accounts of Chemical Research. 46 (4): 927–936. doi:10.1021/ar300083h. PMID 22872015.
- ^ Mahadevi AS, Sastry GN (March 2013). "Cation-π interaction: its role and relevance in chemistry, biology, and material science". Chemical Reviews. 113 (3): 2100–2138. doi:10.1021/cr300222d. PMID 23145968.
- ^ Quiñonero D, Garau C, Rotger C, Frontera A, Ballester P, Costa A, Deyà PM (September 2002). "Anion-pi Interactions: do they exist?". Angewandte Chemie. 41 (18): 3389–3392. doi:10.1002/1521-3773(20020916)41:18<3389::AID-ANIE3389>3.0.CO;2-S. PMID 12298041.
- ^ a b IUPAC (2009). "Hydrophobic interaction". Compendium of Chemical Terminology. doi:10.1351/goldbook.H02907. ISBN 978-0-9678550-9-7. Retrieved 11 November 2013.
- ^ Kronberg B (2016). "The hydrophobic effect". Curr. Opinion Coll. Interface Sci. 22: 14–22. doi:10.1016/j.cocis.2016.02.001.
- ^ Hillyer MB, Gibb BC (May 2016). "Molecular Shape and the Hydrophobic Effect". Annual Review of Physical Chemistry. 67: 307–329. Bibcode:2016ARPC...67..307H. doi:10.1146/annurev-physchem-040215-112316. PMC 5571648. PMID 27215816.
- ^ Ben-Amotz D (May 2016). "Water-Mediated Hydrophobic Interactions". Annual Review of Physical Chemistry. 67: 617–638. Bibcode:2016ARPC...67..617B. doi:10.1146/annurev-physchem-040215-112412. PMID 27215821.
- ^ Snyder PW, Lockett MR, Moustakas DT, Whitesides GM (2014). "Is it the shape of the cavity, or the shape of the water in the cavity?". The European Physical Journal Special Topics. 223 (5): 853–891. Bibcode:2014EPJST.223..853S. doi:10.1140/epjst/e2013-01818-y. S2CID 17572997.
- ^ Biedermann F, Nau WM, Schneider HJ (October 2014). "The hydrophobic effect revisited--studies with supramolecular complexes imply high-energy water as a noncovalent driving force". Angewandte Chemie. 53 (42): 11158–11171. doi:10.1002/anie.201310958. PMID 25070083.
- ^ "Biomolecules: Enzymes". ChemPages Netorials. University of Wisconsin - Madison. Retrieved 27 October 2013.
- ^ Cardo L, Hannon MJ (February 2018). Sigel A, Sigel H, Freisinger E, Sigel RK (eds.). "Non-covalent Metallo-Drugs: Using Shape to Target DNA and RNA Junctions and Other Nucleic Acid Structures". Metal Ions in Life Sciences. 18. de Gruyter GmbH: 303–324. doi:10.1515/9783110470734-017. ISBN 978-3-11-047073-4. PMID 29394030.
- ^ Voet D, Voet JG (2010). Biochemistry (4th ed.). Hoboken, NJ: John Wiley & Sons. ISBN 978-0-470-57095-1.
- ^ Silverman RB (2004). The organic chemistry of drug design and drug action (2. ed.). Amsterdam [u.a.]: Elsevier. ISBN 978-0-12-643732-4.




